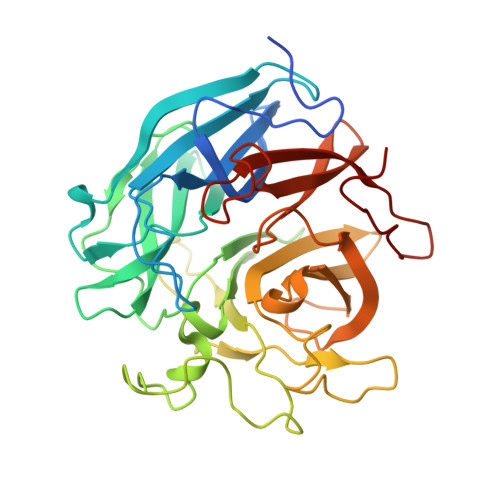
Crystal structure of metagenomic beta-xylosidase/ alpha-l-arabinofuranosidase activated by calcium.
Matsuzawa, T., Kaneko, S., Kishine, N., Fujimoto, Z., Yaoi, K.(2017) J Biochem 162: 173-181
- PubMed: 28204531 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvx012
- Primary Citation Related Structures:
5GLK, 5GLL, 5GLM, 5GLN, 5GLO, 5GLP, 5GLQ, 5GLR - PubMed Abstract:
The crystal structure of metagenomic β-xylosidase/α-l-arabinofuranosidase CoXyl43, activated by calcium ions, was determined in its apo and complexed forms with xylotriose or l-arabinose in the presence and absence of calcium. The presence of calcium ions dramatically increases the kcat of CoXyl43 for p-nitrophenyl β-d-xylopyranoside and reduces the Michaelis constant for p-nitrophenyl α-l-arabinofuranoside. CoXyl43 consists of a single catalytic domain comprised of a five-bladed β-propeller. In the presence of calcium, a single calcium ion was observed at the centre of this catalytic domain, behind the catalytic pocket. In the absence of calcium, the calcium ion was replaced with one sodium ion and one water molecule, and the positions of these cations were shifted by 1.3 Å. The histidine-319 side chain, which coordinates to the 2-hydroxyl oxygen atom of the bound xylose molecule in the catalytic pocket, also coordinates to the calcium ion, but not to the sodium ion. The calcium-dependent increase in activity appears to be caused by the structural change in the catalytic pocket induced by the tightly bound calcium ion and coordinating water molecules, and by the protonation state of glutamic acid-268, the catalytic acid of the enzyme. Our findings further elucidate the complex relationship between metal ions and glycosidases.
- Bioproduction Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), Tsukuba Central 6, 1-1-1 Higashi, Tsukuba, Ibaraki 305-8566, Japan.
Organizational Affiliation: