Structure of glutathionylated hHsp70 SBD (385-641)
Gong, W.B., Yang, J., Zhang, H., Perrett, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Heat shock 70 kDa protein 1A | 257 | Homo sapiens | Mutation(s): 0 Gene Names: HSPA1A, HSP72, HSPA1, HSX70 | 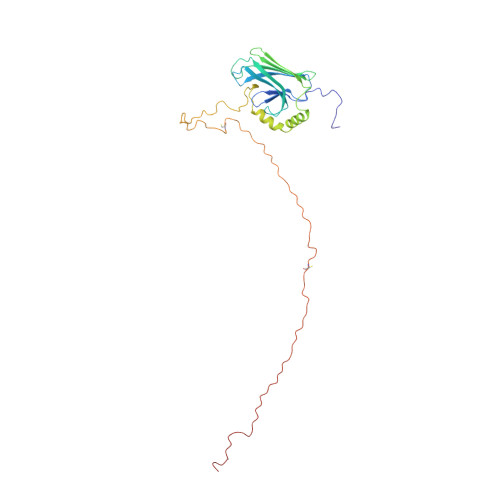 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P0DMV8 GTEx: ENSG00000204389 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DMV8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| SMC Query on SMC | A | L-PEPTIDE LINKING | C4 H9 N O2 S |  | CYS |