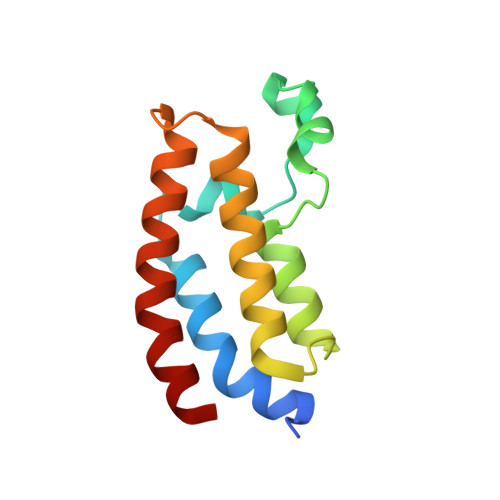
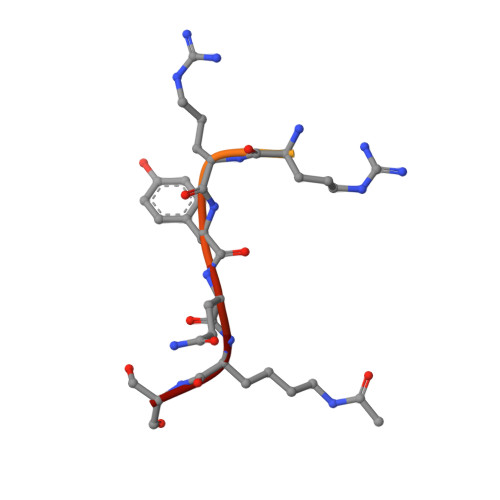
Structural insight into CBP bromodomain-mediated recognition of acetylated histone H3K56ac
Xu, L., Cheng, A., Huang, M., Zhang, J., Jiang, Y., Wang, C., Li, F., Bao, H., Gao, J., Wang, N., Liu, J., Wu, J., Wong, C.C.L., Ruan, K.(2017) FEBS J
- PubMed: 28815970 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14198
- Primary Citation Related Structures:
5GH9 - PubMed Abstract:
The acetylation of lysine 56 of histone H3 (H3K56ac) enhances the binding affinity of histone chaperones to H3-H4 dimers. CREB-binding protein (CBP) possesses a bromodomain that recognizes H3K56 acetylation. CBP also possesses a histone acetyltransferase (HAT) domain, which has been shown to promote H3K56 acetylation of free histones to facilitate delivery of replication-dependent chaperones to acetylated histones for chromatin assembly. However, the mechanism by which the CBP bromodomain recognizes H3K56ac and the context in which such recognition occurs remain elusive. Here, we solved the crystal structure of the CBP bromodomain in complex with an H3K56ac peptide. Our data demonstrate that the CBP bromodomain recognizes H3K56ac with high affinity. Structural and affinity analyses reveal that the CBP bromodomain prefers an aromatic residue at the -2 position and an arginine at the -4 position from the acetyl-lysine, and that the CBP bromodomain selectively recognizes an extended conformation of the H3 αN helix that contains H3K56ac. We also demonstrate that the CBP bromodomain binds to H3K56ac in a recombinant H3-H4 dimer but not in a mono-nucleosome. Our results suggest that the CBP bromodomain selectively recognizes an extended conformation of the K56-acetylated H3 α N region within an H3-H4 dimer, which is expected to facilitate the HAT activity of CBP for subsequent H3K56 acetylation of free histones. Coordinates of the CBP bromodomain in complex with H3K56ac as described in this article have been deposited in the PDB with accession number 5GH9.
- Hefei National Laboratory for Physical Science at the Microscale, School of Life Sciences, University of Science and Technology of China, Hefei, China.
Organizational Affiliation: