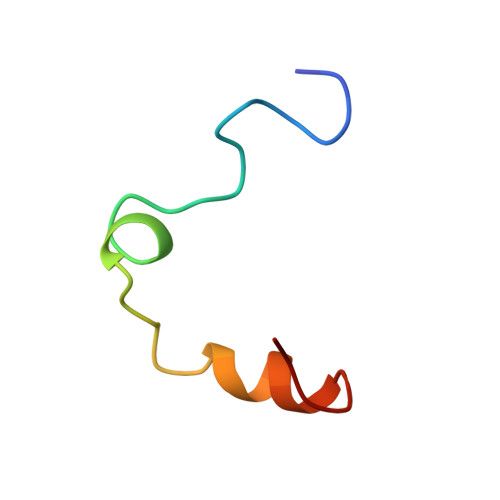
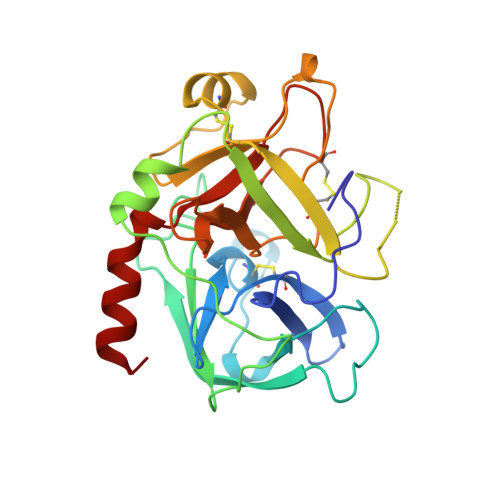
Hirunorms are true hirudin mimetics. The crystal structure of human alpha-thrombin-hirunorm V complex.
De Simone, G., Lombardi, A., Galdiero, S., Nastri, F., Della Morte, R., Staiano, N., Pedone, C., Bolognesi, M., Pavone, V.(1998) Protein Sci 7: 243-253
- PubMed: 9521099 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560070203
- Primary Citation Related Structures:
5GDS - PubMed Abstract:
A novel class of synthetic, multisite-directed thrombin inhibitors, known as hirunorms, has been described recently. These compounds were designed to mimic the binding mode of hirudin, and they have been proven to be very strong and selective thrombin inhibitors. Here we report the crystal structure of the complex formed by human alpha-thrombin and hirunorm V, a 26-residue polypeptide containing non-natural amino acids, determined at 2.1 A resolution and refined to an R-factor of 0.176. The structure reveals that the inhibitor binding mode is distinctive of a true hirudin mimetic, and it highlights the molecular basis of the high inhibitory potency (Ki is in the picomolar range) and the strong selectivity of hirunorm V. Hirunorm V interacts through the N-terminal tetrapeptide with the thrombin active site in a nonsubstrate mode; at the same time, this inhibitor specifically binds through the C-terminal segment to the fibrinogen recognition exosite. The backbone of the N-terminal tetrapeptide Chg1"-Val2"-2-Nal3"-Thr4" (Chg, cyclohexyl-glycine; 2-Nal, beta-(2-naphthyl)-alanine) forms a short beta-strand parallel to thrombin main-chain residues Ser214-Gly219. The Chg1" side chain fills the S2 subsite, Val2" is located at the entrance of S1, whereas 2-Nal3" side chain occupies the aryl-binding site. Such backbone orientation is very close to that observed for the N-terminal residues of hirudin, and it is similar to that of the synthetic retro-binding peptide BMS-183507, but it is opposite to the proposed binding mode of fibrinogen and of small synthetic substrates. Hirunorm V C-terminal segment binds to the fibrinogen recognition exosite, similarly to what observed for hirudin C-termninal tail and related compounds. The linker polypeptide segment connecting hirunorm V N-and C-terminal regions is not observable in the electron density maps. The crystallographic analysis proves the correctness of the design and it provides a compelling proof on the interaction mechanism for this novel class of high potency multisite-directed synthetic thrombin inhibitors.
- Centro Interuniversitario di Ricerca su Peptidi Bioattivi, & Centro di Studio di Biocristallografia-CNR, University of Napoli Federico II, Italy.
Organizational Affiliation: