Resolution and Probabilistic Models of Components in CryoEM Maps of Mature P22 Bacteriophage.
Pintilie, G., Chen, D.H., Haase-Pettingell, C.A., King, J.A., Chiu, W.(2016) Biophys J 110: 827-839
- PubMed: 26743049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2015.11.3522
- Primary Citation Related Structures:
5GAI - PubMed Abstract:
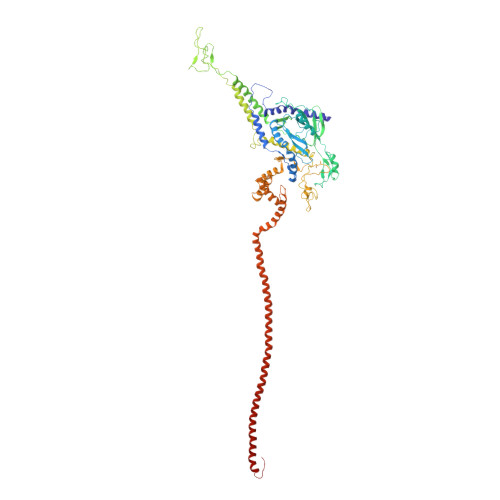
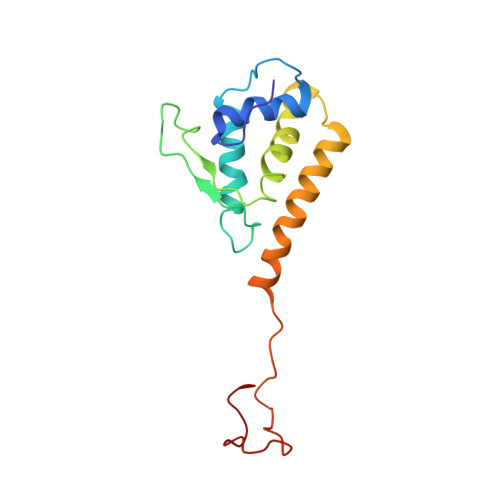
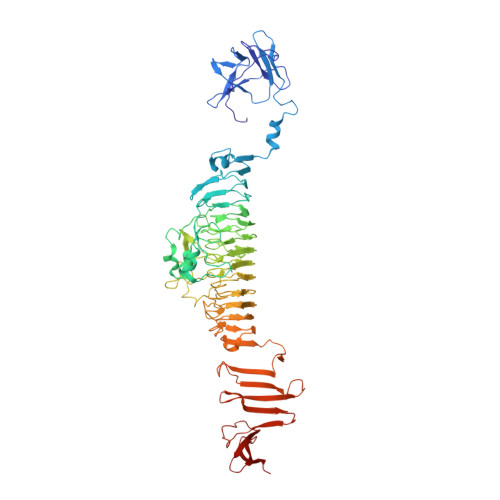
CryoEM continues to produce density maps of larger and more complex assemblies with multiple protein components of mixed symmetries. Resolution is not always uniform throughout a cryoEM map, and it can be useful to estimate the resolution in specific molecular components of a large assembly. In this study, we present procedures to 1) estimate the resolution in subcomponents by gold-standard Fourier shell correlation (FSC); 2) validate modeling procedures, particularly at medium resolutions, which can include loop modeling and flexible fitting; and 3) build probabilistic models that combine high-accuracy priors (such as crystallographic structures) with medium-resolution cryoEM densities. As an example, we apply these methods to new cryoEM maps of the mature bacteriophage P22, reconstructed without imposing icosahedral symmetry. Resolution estimates based on gold-standard FSC show the highest resolution in the coat region (7.6 Å), whereas other components are at slightly lower resolutions: portal (9.2 Å), hub (8.5 Å), tailspike (10.9 Å), and needle (10.5 Å). These differences are indicative of inherent structural heterogeneity and/or reconstruction accuracy in different subcomponents of the map. Probabilistic models for these subcomponents provide new insights, to our knowledge, and structural information when taking into account uncertainty given the limitations of the observed density.
- Verna and Marrs McLean Department of Biochemistry and Molecular Biology, Baylor College of Medicine, Houston, Texas. Electronic address: pintilie@bcm.edu.
Organizational Affiliation: