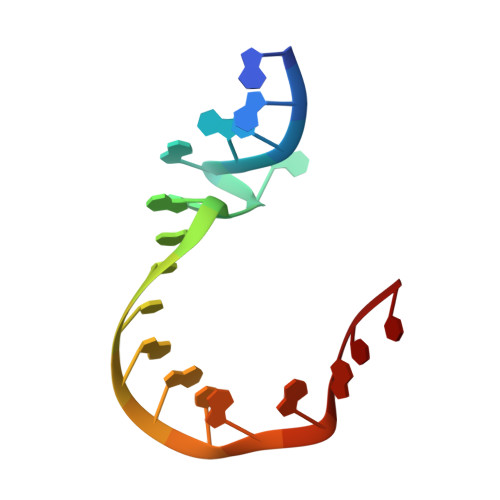
A Quasi-Cyclic RNA Nano-Scale Molecular Object Constructed Using Kink Turns.
Huang, L., Lilley, D.M.J.(2016) Nanoscale 8: 15189
- PubMed: 27506301 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c6nr05186c
- Primary Citation Related Structures:
5G4T, 5G4U, 5G4V - PubMed Abstract:
k-Turns are widespread RNA architectural elements that mediate tertiary interactions. We describe a double-kink-turn motif comprising two inverted k-turns that forms a tight horse-shoe structure that can assemble into a variety of shapes by coaxial association of helical ends. Using X-ray crystallography we show that these assemble with two (dumbell), three (triangle) and four units (square), with or without bound protein, within the crystal lattice. In addition, exchange of a single basepair can almost double the pore radius or shape of a molecular assembly. On the basis of this analysis we synthesized a 114 nt self-complementary RNA containing six k-turns. The crystal structure of this species shows that it forms a quasi-cyclic triangular object. These are randomly disposed about the three-fold axis in the crystal lattice, generating a circular RNA of quasi D3 symmetry with a shape reminiscent of that of a cyclohexane molecule in its chair conformation. This work demonstrates that the k-turn is a powerful building block in the construction of nano-scale molecular objects, and illustrates why k-turns are widely used in natural RNA molecules to organize long-range architecture and mediate tertiary contacts.
- Cancer Research UK Nucleic Acid Structure Research Group, MSI/WTB Complex, The University of Dundee, Dow Street, Dundee DD1 5EH, UK. d.m.j.lilley@dundee.ac.uk l.y.huang@dundee.ac.uk.
Organizational Affiliation: