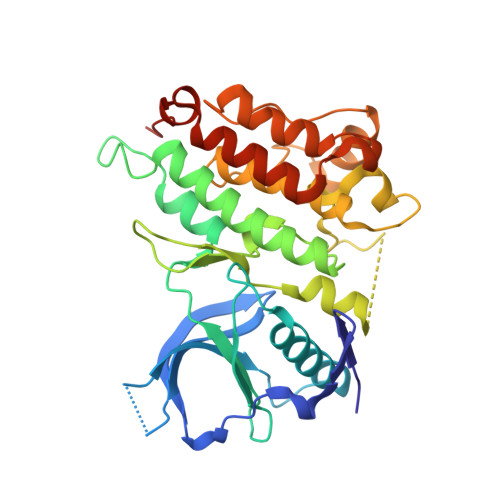
Discovery of Entrectinib: A New 3-Aminoindazole as a Potent Anaplastic Lymphoma Kinase (Alk), C-Ros Oncogene 1 Kinase (Ros1), and Pan-Tropomyosin Receptor Kinases (Pan-Trks) Inhibitor.
Menichincheri, M., Ardini, E., Magnaghi, P., Avanzi, N., Banfi, P., Bossi, R.T., Buffa, L., Canevari, G., Ceriani, L., Colombo, M., Corti, L., Donati, D., Fasolini, M., Felder, E.R., Fiorelli, C., Fiorentini, F., Galvani, A., Isacchi, A., Borgia, A.L., Marchionni, C., Nesi, M., Orrenius, C., Panzeri, A., Pesenti, E., Rusconi, L., Saccardo, B.M., Vanotti, E., Perrone, E., Orsini, P.(2016) J Med Chem 59: 3392
- PubMed: 27003761 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00064
- Primary Citation Related Structures:
5FTO, 5FTQ - PubMed Abstract:
Anaplastic lymphoma kinase (ALK) is a receptor tyrosine kinase responsible for the development of different tumor types. Despite the remarkable clinical activity of crizotinib (Xalkori), the first ALK inhibitor approved in 2011, the emergence of resistance mutations and of brain metastases frequently causes relapse in patients. Within our ALK drug discovery program, we identified compound 1, a novel 3-aminoindazole active on ALK in biochemical and in cellular assays. Its optimization led to compound 2 (entrectinib), a potent orally available ALK inhibitor active on ALK-dependent cell lines, efficiently penetrant the blood-brain barrier (BBB) in different animal species and highly efficacious in in vivo xenograft models. Moreover, entrectinib resulted to be strictly potent on the closely related tyrosine kinases ROS1 and TRKs recently found constitutively activated in several tumor types. Entrectinib is currently undergoing phase I/II clinical trial for the treatment of patients affected by ALK-, ROS1-, and TRK-positive tumors.
- Oncology, Nerviano Medical Sciences Srl , Viale Pasteur 10, 20014 Nerviano, Milan, Italy.
Organizational Affiliation: