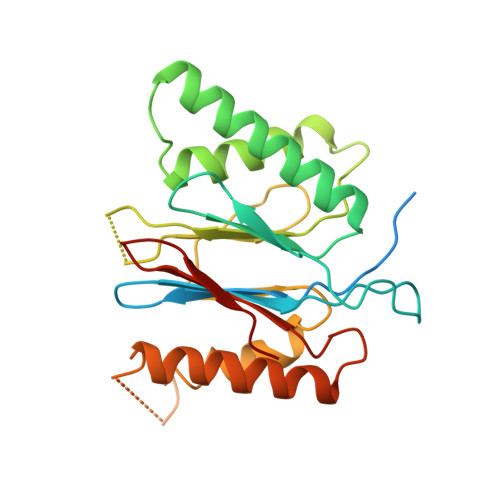
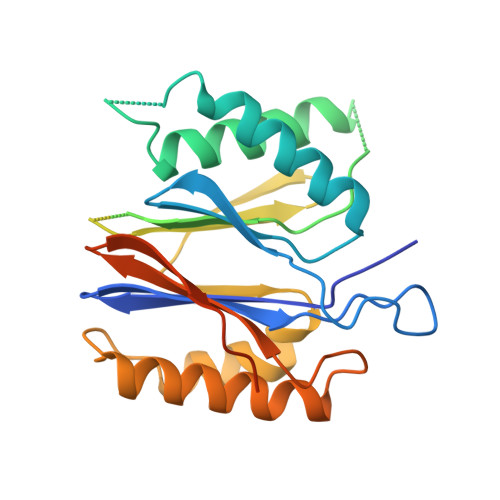
Structure and Function Based Design of Plasmodium-Selective Proteasome Inhibitors
Li, H., O'Donoghue, A.J., Van Der Linden, W.A., Xie, S.C., Yoo, E., Foe, I.T., Tilley, L., Craik, C.S., Da Fonseca, P.C.A., Bogyo, M.(2016) Nature 530: 233
- PubMed: 26863983 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature16936
- Primary Citation Related Structures:
5FMG - PubMed Abstract:
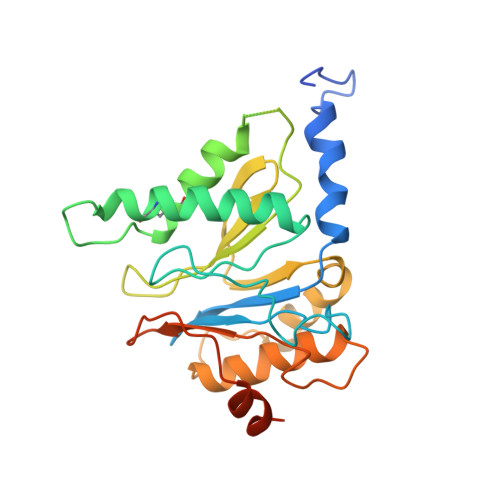
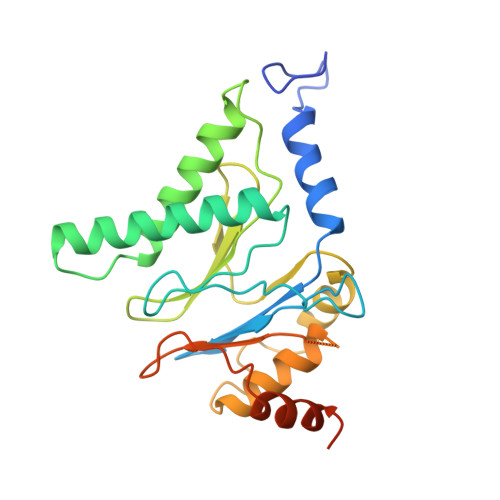
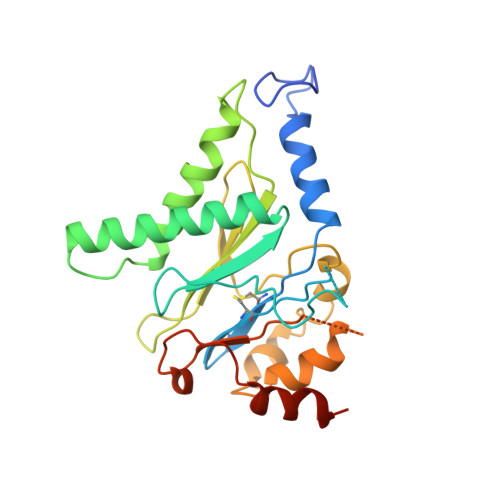
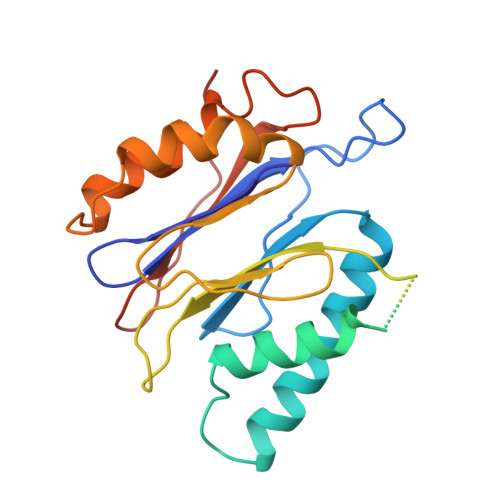
The proteasome is a multi-component protease complex responsible for regulating key processes such as the cell cycle and antigen presentation. Compounds that target the proteasome are potentially valuable tools for the treatment of pathogens that depend on proteasome function for survival and replication. In particular, proteasome inhibitors have been shown to be toxic for the malaria parasite Plasmodium falciparum at all stages of its life cycle. Most compounds that have been tested against the parasite also inhibit the mammalian proteasome, resulting in toxicity that precludes their use as therapeutic agents. Therefore, better definition of the substrate specificity and structural properties of the Plasmodium proteasome could enable the development of compounds with sufficient selectivity to allow their use as anti-malarial agents. To accomplish this goal, here we use a substrate profiling method to uncover differences in the specificities of the human and P. falciparum proteasome. We design inhibitors based on amino-acid preferences specific to the parasite proteasome, and find that they preferentially inhibit the β2-subunit. We determine the structure of the P. falciparum 20S proteasome bound to the inhibitor using cryo-electron microscopy and single-particle analysis, to a resolution of 3.6 Å. These data reveal the unusually open P. falciparum β2 active site and provide valuable information about active-site architecture that can be used to further refine inhibitor design. Furthermore, consistent with the recent finding that the proteasome is important for stress pathways associated with resistance of artemisinin family anti-malarials, we observe growth inhibition synergism with low doses of this β2-selective inhibitor in artemisinin-sensitive and -resistant parasites. Finally, we demonstrate that a parasite-selective inhibitor could be used to attenuate parasite growth in vivo without appreciable toxicity to the host. Thus, the Plasmodium proteasome is a chemically tractable target that could be exploited by next-generation anti-malarial agents.
- Department of Chemical and Systems Biology, Stanford University School of Medicine, Stanford, California 94305, USA.
Organizational Affiliation: