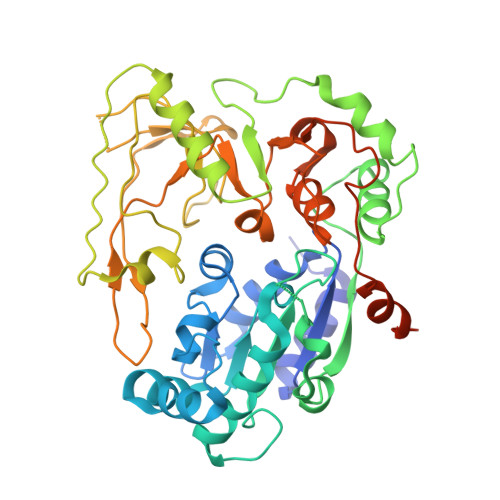
Structural and Functional Insights into the Unwinding Mechanism of Bacteroides sp Pif1
Zhou, X., Ren, W., Bharath, S.R., Tang, X., He, Y., Chen, C., Liu, Z., Li, D., Song, H.(2016) Cell Rep 14: 2030-2039
- PubMed: 26904952 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2016.02.008
- Primary Citation Related Structures:
5FHD, 5FHE, 5FHF, 5FHG, 5FHH - PubMed Abstract:
Pif1 is a conserved SF1B DNA helicase involved in maintaining genome stability through unwinding double-stranded DNAs (dsDNAs), DNA/RNA hybrids, and G quadruplex (G4) structures. Here, we report the structures of the helicase domain of human Pif1 and Bacteroides sp Pif1 (BaPif1) in complex with ADP-AlF4(-) and two different single-stranded DNAs (ssDNAs). The wedge region equivalent to the β hairpin in other SF1B DNA helicases folds into an extended loop followed by an α helix. The Pif1 signature motif of BaPif1 interacts with the wedge region and a short helix in order to stabilize these ssDNA binding elements, therefore indirectly exerting its functional role. Domain 2B of BaPif1 undergoes a large conformational change upon concomitant binding of ATP and ssDNA, which is critical for Pif1's activities. BaPif1 cocrystallized with a tailed dsDNA and ADP-AlF4(-), resulting in a bound ssDNA bent nearly 90° at the ssDNA/dsDNA junction. The conformational snapshots of BaPif1 provide insights into the mechanism governing the helicase activity of Pif1.
- Life Sciences Institute and Innovation Center for Cell Signaling Network, Zhejiang University, 388 Yuhangtang Road, Hangzhou 310058, China; Institute of Molecular and Cell Biology, 61 Biopolis Drive, Singapore 138673, Singapore.
Organizational Affiliation: