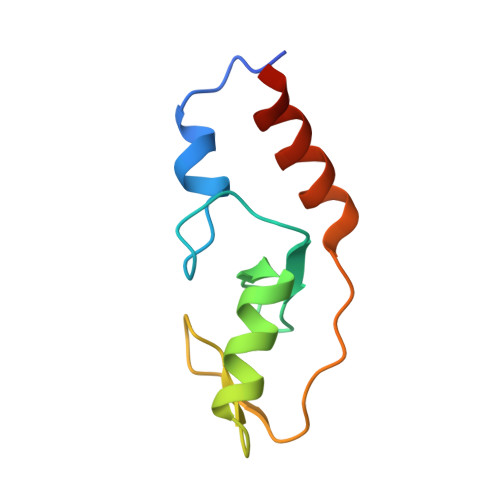
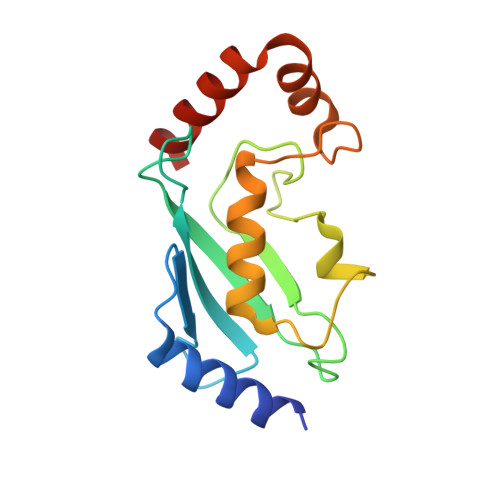
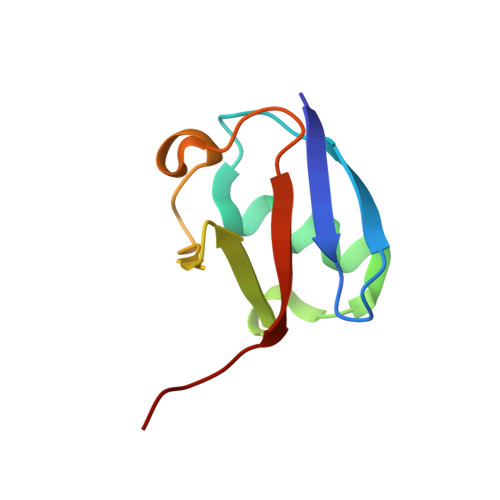
Functional role of TRIM E3 ligase oligomerization and regulation of catalytic activity.
Koliopoulos, M.G., Esposito, D., Christodoulou, E., Taylor, I.A., Rittinger, K.(2016) EMBO J 35: 1204-1218
- PubMed: 27154206 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.201593741
- Primary Citation Related Structures:
5FER, 5FEY - PubMed Abstract:
TRIM E3 ubiquitin ligases regulate a wide variety of cellular processes and are particularly important during innate immune signalling events. They are characterized by a conserved tripartite motif in their N-terminal portion which comprises a canonical RING domain, one or two B-box domains and a coiled-coil region that mediates ligase dimerization. Self-association via the coiled-coil has been suggested to be crucial for catalytic activity of TRIMs; however, the precise molecular mechanism underlying this observation remains elusive. Here, we provide a detailed characterization of the TRIM ligases TRIM25 and TRIM32 and show how their oligomeric state is linked to catalytic activity. The crystal structure of a complex between the TRIM25 RING domain and an ubiquitin-loaded E2 identifies the structural and mechanistic features that promote a closed E2~Ub conformation to activate the thioester for ubiquitin transfer allowing us to propose a model for the regulation of activity in the full-length protein. Our data reveal an unexpected diversity in the self-association mechanism of TRIMs that might be crucial for their biological function.
- Mill Hill Laboratory, Molecular Structure of Cell Signalling Laboratory, The Francis Crick Institute, London, UK.
Organizational Affiliation: