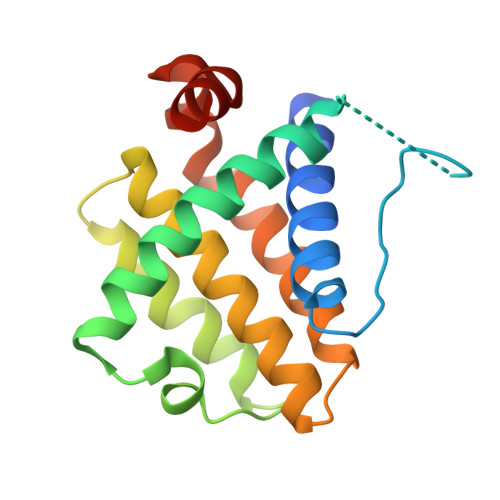
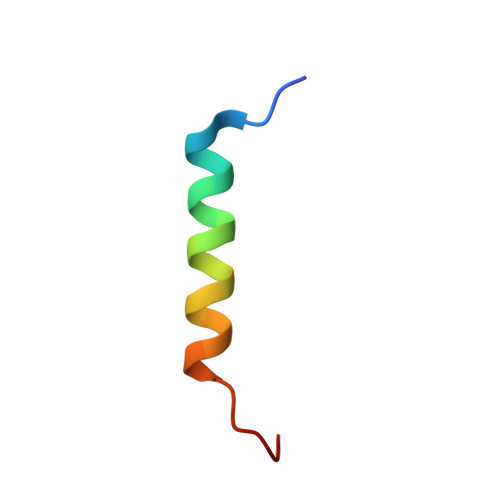
Structural and biochemical analysis of Bcl-2 interaction with the hepatitis B virus protein HBx
Jiang, T.Y., Liu, M.H., Wu, J.P., Shi, Y.G.(2016) Proc Natl Acad Sci U S A
- PubMed: 26858413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1525616113
- Primary Citation Related Structures:
5FCG - PubMed Abstract:
HBx is a hepatitis B virus protein that is required for viral infectivity and replication. Anti-apoptotic Bcl-2 family members are thought to be among the important host targets of HBx. However, the structure and function of HBx are poorly understood and the molecular mechanism of HBx-induced carcinogenesis remains unknown. In this study, we report biochemical and structural characterization of HBx. The recombinant HBx protein contains metal ions, in particular iron and zinc. A BH3-like motif in HBx (residues 110-135) binds Bcl-2 with a dissociation constant of ∼193 μM, which is drastically lower than that for a canonical BH3 motif from Bim or Bad. Structural analysis reveals that, similar to other BH3 motifs, the BH3-like motif of HBx adopts an amphipathic α-helix and binds the conserved BH3-binding groove on Bcl-2. Unlike the helical Bim or Bad BH3 motif, the C-terminal portion of the bound HBx BH3-like motif has an extended conformation and makes considerably fewer interactions with Bcl-2. These observations suggest that HBx may modulate Bcl-2 function in a way that is different from that of the classical BH3-only proteins.
- Ministry of Education Key Laboratory of Protein Science, Tsinghua-Peking Joint Center for Life Sciences, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: