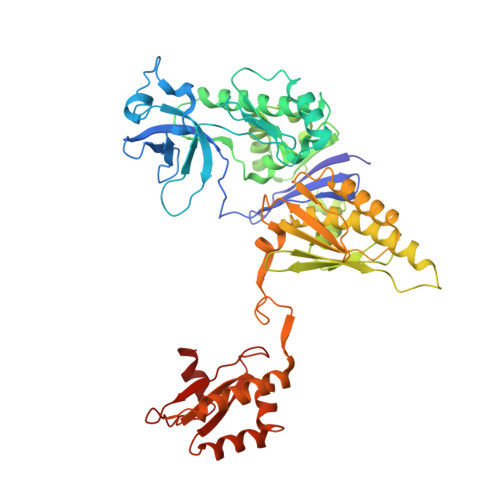
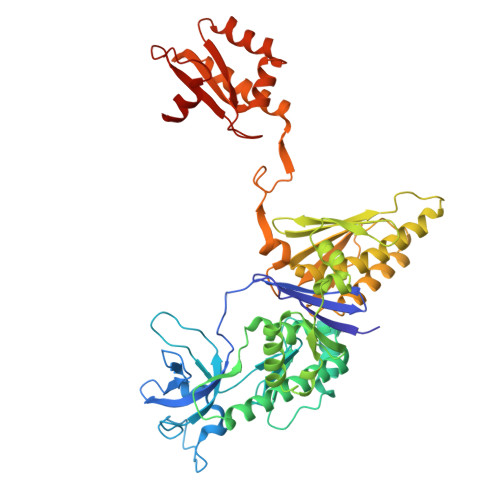
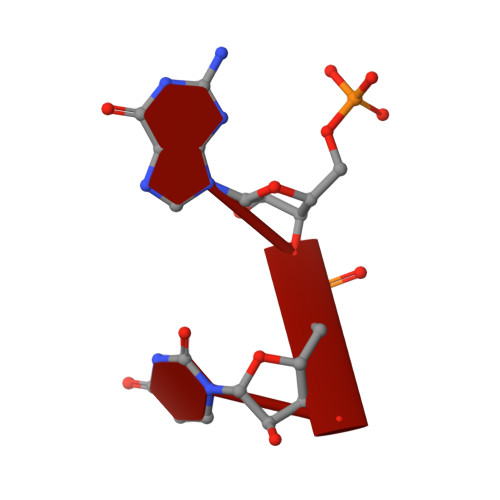
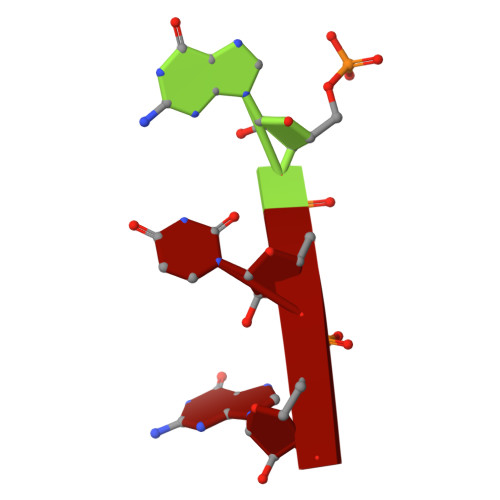
Substrate Recognition and Autoinhibition in the Central Ribonuclease RNase E.
Bandyra, K.J., Wandzik, J.M., Luisi, B.F.(2018) Mol Cell 72: 275-285.e4
- PubMed: 30270108 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2018.08.039
- Primary Citation Related Structures:
5F6C, 6G63 - PubMed Abstract:
The endoribonuclease RNase E is a principal factor in RNA turnover and processing that helps to exercise fine control of gene expression in bacteria. While its catalytic activity can be strongly influenced by the chemical identity of the 5' end of RNA substrates, the enzyme can also cleave numerous substrates irrespective of the chemistry of their 5' ends through a mechanism that has remained largely unexplained. We report structural and functional data illuminating details of both operational modes. Our crystal structure of RNase E in complex with the sRNA RprA reveals a duplex recognition site that saddles an inter-protomer surface to help present substrates for cleavage. Our data also reveal an autoinhibitory pocket that modulates the overall activity of the ribonuclease. Taking these findings together, we propose how RNase E uses versatile modes of RNA recognition to achieve optimal activity and specificity.
- Department of Biochemistry, University of Cambridge, Tennis Court Road, Cambridge CB2 1GA, UK. Electronic address: kjb69@cam.ac.uk.
Organizational Affiliation: