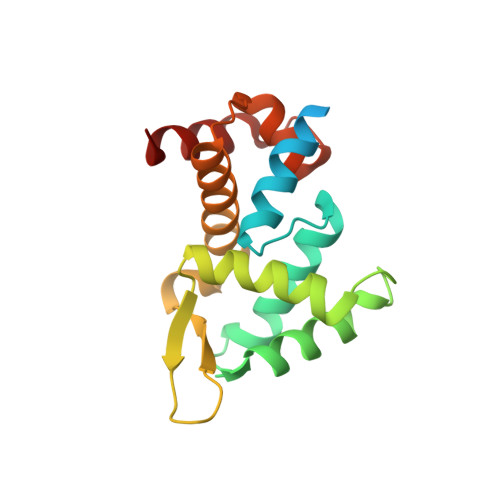
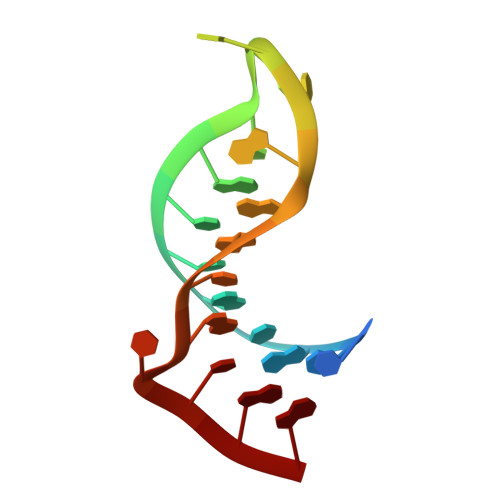
Roquin recognizes a non-canonical hexaloop structure in the 3'-UTR of Ox40.
Janowski, R., Heinz, G.A., Schlundt, A., Wommelsdorf, N., Brenner, S., Gruber, A.R., Blank, M., Buch, T., Buhmann, R., Zavolan, M., Niessing, D., Heissmeyer, V., Sattler, M.(2016) Nat Commun 7: 11032-11032
- PubMed: 27010430 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11032
- Primary Citation Related Structures:
5F5F, 5F5H - PubMed Abstract:
The RNA-binding protein Roquin is required to prevent autoimmunity. Roquin controls T-helper cell activation and differentiation by limiting the induced expression of costimulatory receptors such as tumor necrosis factor receptor superfamily 4 (Tnfrs4 or Ox40). A constitutive decay element (CDE) with a characteristic triloop hairpin was previously shown to be recognized by Roquin. Here we use SELEX assays to identify a novel U-rich hexaloop motif, representing an alternative decay element (ADE). Crystal structures and NMR data show that the Roquin-1 ROQ domain recognizes hexaloops in the SELEX-derived ADE and in an ADE-like variant present in the Ox40 3'-UTR with identical binding modes. In cells, ADE-like and CDE-like motifs cooperate in the repression of Ox40 by Roquin. Our data reveal an unexpected recognition of hexaloop cis elements for the posttranscriptional regulation of target messenger RNAs by Roquin.
- Group Intracellular Transport and RNA Biology, Institute of Structural Biology, Helmholtz Zentrum München, Ingolstädter Landstrasse 1, Neuherberg DE-85764, Germany.
Organizational Affiliation: