Crystal structure of anti-HCV E2 antibody HC84-26
Gao, M., Mariuzza, R.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Anti-HCV E2 glycoprotein Fab heavy chain | 217 | Homo sapiens | Mutation(s): 0 | 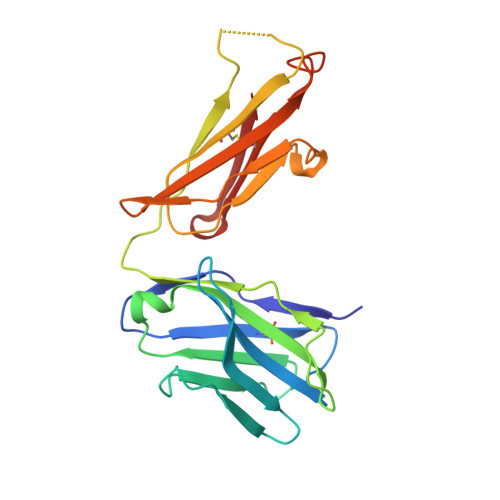 | |
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Anti-HCV E2 glycoprotein Fab light chain | 214 | Homo sapiens | Mutation(s): 0 | 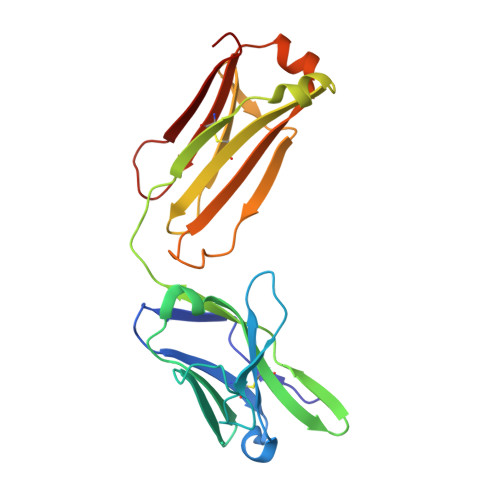 | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 64.736 | α = 90 |
| b = 69.806 | β = 90 |
| c = 87.829 | γ = 90 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data collection |
| HKL-2000 | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| Coot | model building |
| PDB_EXTRACT | data extraction |