Potent, selective, and orally bioavailable inhibitors of VPS34 provide chemical tools to modulate autophagy in vivo
Kearney, E.P.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Phosphatidylinositol 3-kinase catalytic subunit type 3 | 625 | Homo sapiens | Mutation(s): 0 Gene Names: PIK3C3, VPS34 EC: 2.7.1.137 | 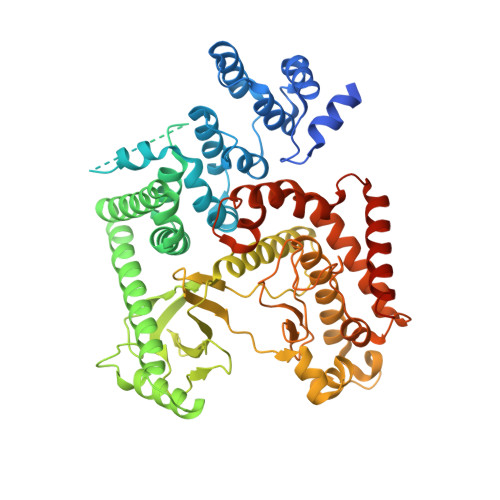 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q8NEB9 GTEx: ENSG00000078142 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8NEB9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 5QS Download:Ideal Coordinates CCD File | C [auth A], E [auth B] | 1-[[4-(cyclopropylmethyl)-5-[2-(pyridin-4-ylamino)pyrimidin-4-yl]pyrimidin-2-yl]amino]-2-methyl-propan-2-ol C21 H25 N7 O XJTIGGCBXFIZJV-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | H [auth B] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | G [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| NA Download:Ideal Coordinates CCD File | D [auth A], F [auth B] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 113.726 | α = 90 |
| b = 114.481 | β = 90 |
| c = 143.903 | γ = 90 |
| Software Name | Purpose |
|---|---|
| BUSTER | refinement |
| Aimless | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| PHASER | phasing |