A Therapeutic Antibody for Cancer, Derived from Single Human B Cells.
Bushey, R.T., Moody, M.A., Nicely, N.L., Haynes, B.F., Alam, S.M., Keir, S.T., Bentley, R.C., Roy Choudhury, K., Gottlin, E.B., Campa, M.J., Liao, H.X., Patz, E.F.(2016) Cell Rep 15: 1505-1513
- PubMed: 27160908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2016.04.038
- Primary Citation Related Structures:
5EA0 - PubMed Abstract:
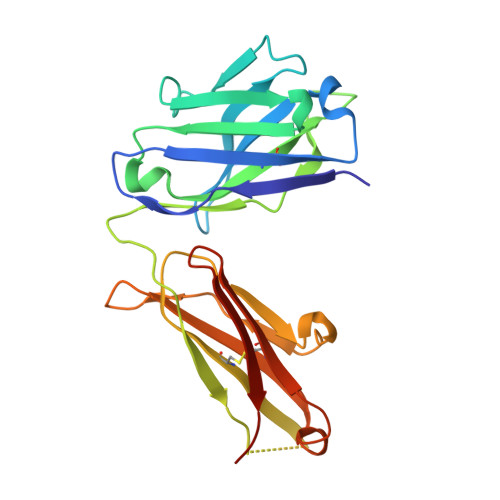
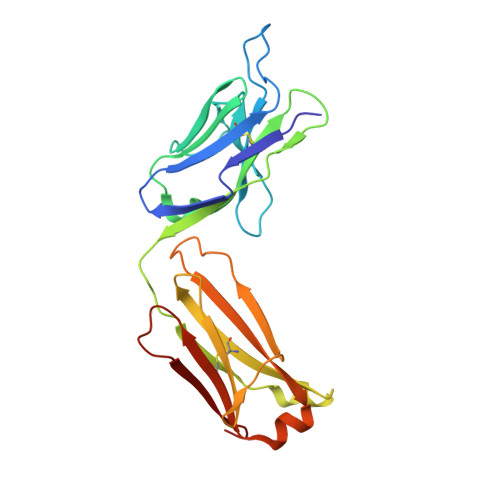
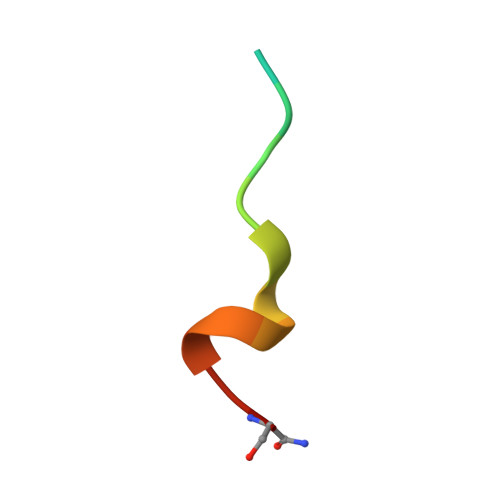
Some patients with cancer never develop metastasis, and their host response might provide cues for innovative treatment strategies. We previously reported an association between autoantibodies against complement factor H (CFH) and early-stage lung cancer. CFH prevents complement-mediated cytotoxicity (CDC) by inhibiting formation of cell-lytic membrane attack complexes on self-surfaces. In an effort to translate these findings into a biologic therapy for cancer, we isolated and expressed DNA sequences encoding high-affinity human CFH antibodies directly from single, sorted B cells obtained from patients with the antibody. The co-crystal structure of a CFH antibody-target complex shows a conformational change in the target relative to the native structure. This recombinant CFH antibody causes complement activation and release of anaphylatoxins, promotes CDC of tumor cell lines, and inhibits tumor growth in vivo. The isolation of anti-tumor antibodies derived from single human B cells represents an alternative paradigm in antibody drug discovery.
- Department of Radiology, Duke University Medical Center, Durham, NC 27710, USA.
Organizational Affiliation: