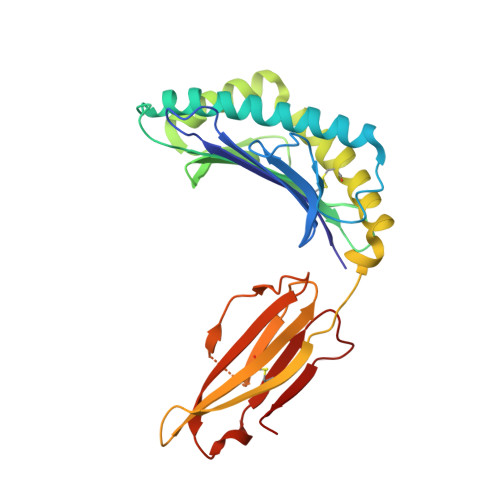
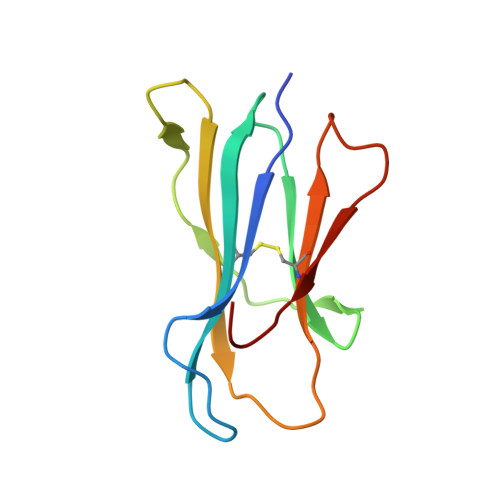
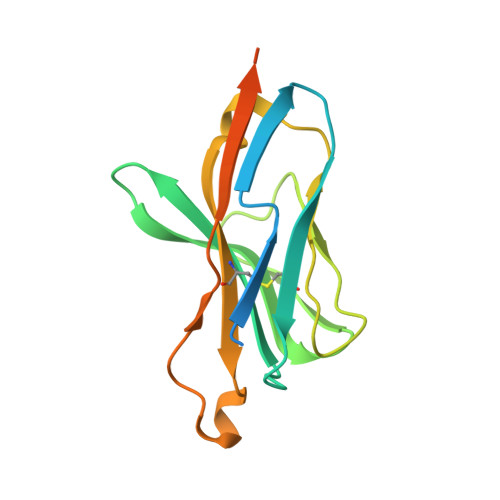
An Engineered Switch in T Cell Receptor Specificity Leads to an Unusual but Functional Binding Geometry.
Harris, D.T., Singh, N.K., Cai, Q., Smith, S.N., Vander Kooi, C.W., Procko, E., Kranz, D.M., Baker, B.M.(2016) Structure 24: 1142-1154
- PubMed: 27238970 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.04.011
- Primary Citation Related Structures:
5E9D - PubMed Abstract:
Utilizing a diverse binding site, T cell receptors (TCRs) specifically recognize a composite ligand comprised of a foreign peptide and a major histocompatibility complex protein (MHC). To help understand the determinants of TCR specificity, we studied a parental and engineered receptor whose peptide specificity had been switched via molecular evolution. Altered specificity was associated with a significant change in TCR-binding geometry, but this did not impact the ability of the TCR to signal in an antigen-specific manner. The determinants of binding and specificity were distributed among contact and non-contact residues in germline and hypervariable loops, and included disruption of key TCR-MHC interactions that bias αβ TCRs toward particular binding modes. Sequence-fitness landscapes identified additional mutations that further enhanced specificity. Our results demonstrate that TCR specificity arises from the distributed action of numerous sites throughout the interface, with significant implications for engineering therapeutic TCRs with novel and functional recognition properties.
- Department of Biochemistry, University of Illinois, 600 S. Matthews Ave., Urbana, IL 61801, USA.
Organizational Affiliation: