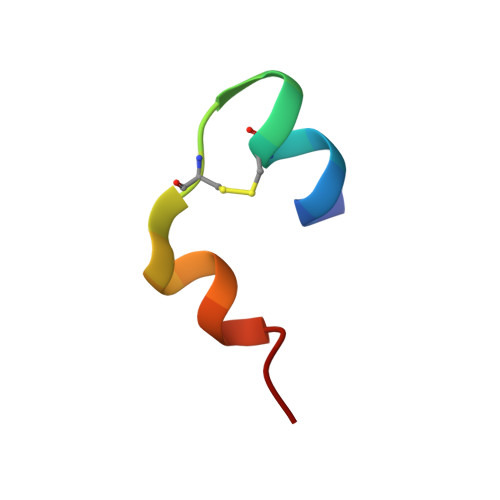
Ultra-high resolution X-ray structures of two forms of human recombinant insulin at 100 K.
Lisgarten, D.R., Palmer, R.A., Lobley, C.M.C., Naylor, C.E., Chowdhry, B.Z., Al-Kurdi, Z.I., Badwan, A.A., Howlin, B.J., Gibbons, N.C.J., Saldanha, J.W., Lisgarten, J.N., Basak, A.K.(2017) Chem Cent J 11: 73-73
- PubMed: 29086855 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s13065-017-0296-y
- Primary Citation Related Structures:
5E7W - PubMed Abstract:
The crystal structure of a commercially available form of human recombinant (HR) insulin, Insugen (I), used in the treatment of diabetes has been determined to 0.92 Å resolution using low temperature, 100 K, synchrotron X-ray data collected at 16,000 keV (λ = 0.77 Å). Refinement carried out with anisotropic displacement parameters, removal of main-chain stereochemical restraints, inclusion of H atoms in calculated positions, and 220 water molecules, converged to a final value of R = 0.1112 and R free = 0.1466. The structure includes what is thought to be an ordered propanol molecule (POL) only in chain D(4) and a solvated acetate molecule (ACT) coordinated to the Zn atom only in chain B(2). Possible origins and consequences of the propanol and acetate molecules are discussed. Three types of amino acid representation in the electron density are examined in detail: (i) sharp with very clearly resolved features; (ii) well resolved but clearly divided into two conformations which are well behaved in the refinement, both having high quality geometry; (iii) poor density and difficult or impossible to model. An example of type (ii) is observed for the intra-chain disulphide bridge in chain C(3) between Sγ6-Sγ11 which has two clear conformations with relative refined occupancies of 0.8 and 0.2, respectively. In contrast the corresponding S-S bridge in chain A(1) shows one clearly defined conformation. A molecular dynamics study has provided a rational explanation of this difference between chains A and C. More generally, differences in the electron density features between corresponding residues in chains A and C and chains B and D is a common observation in the Insugen (I) structure and these effects are discussed in detail. The crystal structure, also at 0.92 Å and 100 K, of a second commercially available form of human recombinant insulin, Intergen (II), deposited in the Protein Data Bank as 3W7Y which remains otherwise unpublished is compared here with the Insugen (I) structure. In the Intergen (II) structure there is no solvated propanol or acetate molecule. The electron density of Intergen (II), however, does also exhibit the three types of amino acid representations as in Insugen (I). These effects do not necessarily correspond between chains A and C or chains B and D in Intergen (II), or between corresponding residues in Insugen (I). The results of this comparison are reported. Graphical abstract Conformations of PheB25 and PheD25 in three insulin structures: implications for biological activity? Insulin residues PheB25 and PheD25 are considered to be important for insulin receptor binding and changes in biological activity occur when these residues are modified. In porcine insulin and Intergen (II) PheB25 adopts conformation B and PheD25 conformation D. However, unexpectedly PheB25 in Insugen (I) human recombinant insulin adopts two distinct conformations corresponding to B and D, Figure 1 and PheD25 adopts a single conformation corresponding to B not D, Figure 2. Conformations of this residue in the ultra-high resolution structure of Insugen (I) are therefore unique within this set. Figures were produced with Biovia, Discovery Studio 2016.
- Biomolecular Research Group, School of Human and Life Sciences, Canterbury Christ Church University, North Holmes Road, Canterbury, Kent, CT1 1QU, UK.
Organizational Affiliation: