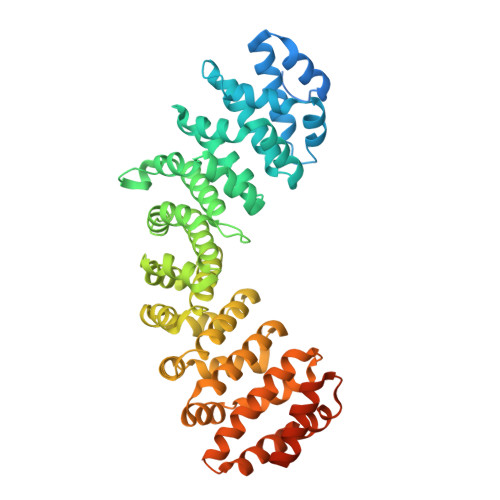
Nuclear Localization of the DNA Repair Scaffold XRCC1: Uncovering the Functional Role of a Bipartite NLS.
Kirby, T.W., Gassman, N.R., Smith, C.E., Pedersen, L.C., Gabel, S.A., Sobhany, M., Wilson, S.H., London, R.E.(2015) Sci Rep 5: 13405-13405
- PubMed: 26304019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep13405
- Primary Citation Related Structures:
5E6Q - PubMed Abstract:
We have characterized the nuclear localization signal (NLS) of XRCC1 structurally using X-ray crystallography and functionally using fluorescence imaging. Crystallography and binding studies confirm the bipartite nature of the XRCC1 NLS interaction with Importin α (Impα) in which the major and minor binding motifs are separated by >20 residues, and resolve previous inconsistent determinations. Binding studies of peptides corresponding to the bipartite NLS, as well as its major and minor binding motifs, to both wild-type and mutated forms of Impα reveal pronounced cooperative binding behavior that is generated by the proximity effect of the tethered major and minor motifs of the NLS. The cooperativity stems from the increased local concentration of the second motif near its cognate binding site that is a consequence of the stepwise binding behavior of the bipartite NLS. We predict that the stepwise dissociation of the NLS from Impα facilitates unloading by providing a partially complexed intermediate that is available for competitive binding by Nup50 or the Importin β binding domain. This behavior provides a basis for meeting the intrinsically conflicting high affinity and high flux requirements of an efficient nuclear transport system.
- Genome Integrity &Structural Biology Laboratory, NIEHS, National Institutes of Health, Research Triangle Park, North Carolina 27709.
Organizational Affiliation: