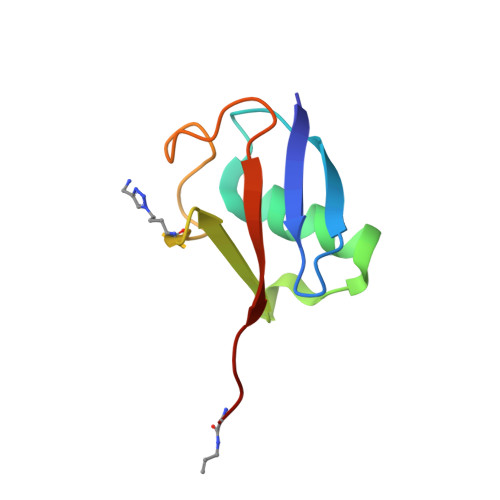
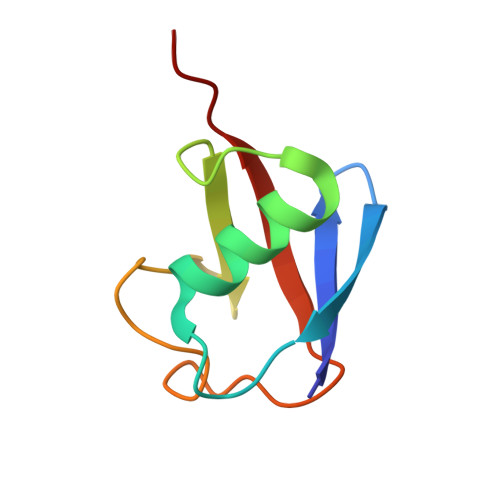
Recognition of Lys48-Linked Di-ubiquitin and Deubiquitinating Activities of the SARS Coronavirus Papain-like Protease.
Bekes, M., van der Heden van Noort, G.J., Ekkebus, R., Ovaa, H., Huang, T.T., Lima, C.D.(2016) Mol Cell 62: 572-585
- PubMed: 27203180 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2016.04.016
- Primary Citation Related Structures:
5E6J - PubMed Abstract:
Deubiquitinating enzymes (DUBs) recognize and cleave linkage-specific polyubiquitin (polyUb) chains, but mechanisms underlying specificity remain elusive in many cases. The severe acute respiratory syndrome (SARS) coronavirus papain-like protease (PLpro) is a DUB that cleaves ISG15, a two-domain Ub-like protein, and Lys48-linked polyUb chains, releasing diUb(Lys48) products. To elucidate this specificity, we report the 2.85 Å crystal structure of SARS PLpro bound to a diUb(Lys48) activity-based probe. SARS PLpro binds diUb(Lys48) in an extended conformation via two contact sites, S1 and S2, which are proximal and distal to the active site, respectively. We show that specificity for polyUb(Lys48) chains is predicated on contacts in the S2 site and enhanced by an S1-S1' preference for a Lys48 linkage across the active site. In contrast, ISG15 specificity is dominated by contacts in the S1 site. Determinants revealed for polyUb(Lys48) specificity should prove useful in understanding PLpro deubiquitinating activities in coronavirus infections.
- Department of Biochemistry and Molecular Pharmacology, New York University School of Medicine, New York, NY 10016, USA; Structural Biology Program, Sloan Kettering Institute, 1275 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: