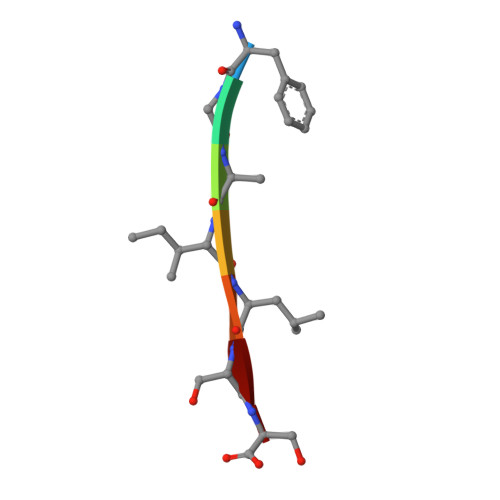
Crystal Structures of IAPP Amyloidogenic Segments Reveal a Novel Packing Motif of Out-of-Register Beta Sheets.
Soriaga, A.B., Sangwan, S., Macdonald, R., Sawaya, M.R., Eisenberg, D.(2016) J Phys Chem B 120: 5810-5816
- PubMed: 26629790 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jpcb.5b09981
- Primary Citation Related Structures:
5E5V, 5E5X, 5E5Z, 5E61 - PubMed Abstract:
Structural studies of amyloidogenic segments by X-ray crystallography have revealed a novel packing motif, consisting of out-of-register β sheets, which may constitute one of the toxic species in aggregation related diseases. Here we sought to determine the presence of such a motif in islet amyloid polypeptide (IAPP), whose amyloidogenic properties are associated with type 2 diabetes. We determined four new crystal structures of segments within IAPP, all forming steric zippers. Most interestingly, one of the segments in the fibril core of IAPP forms an out-of-register steric zipper. Analysis of this structure reveals several commonalities with previously solved out-of-register fibrils. Our results provide additional evidence of out-of-register β sheets as a common structural motif in amyloid aggregates.
- Howard Hughes Medical Institute, ‡UCLA-DOE Institute of Genomics and Proteomics, §Department of Biological Chemistry, and ∥Department of Chemistry & Biochemistry, UCLA , Los Angeles, California 90095, United States.
Organizational Affiliation: