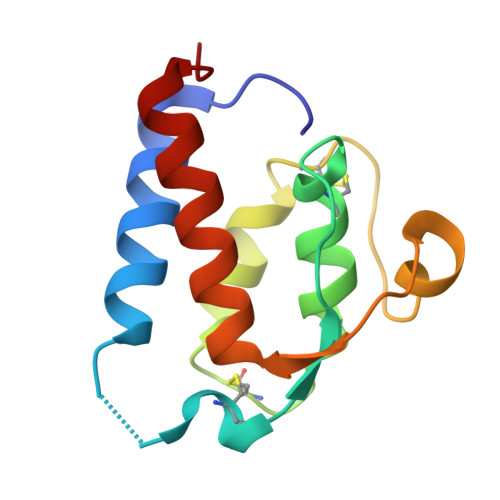
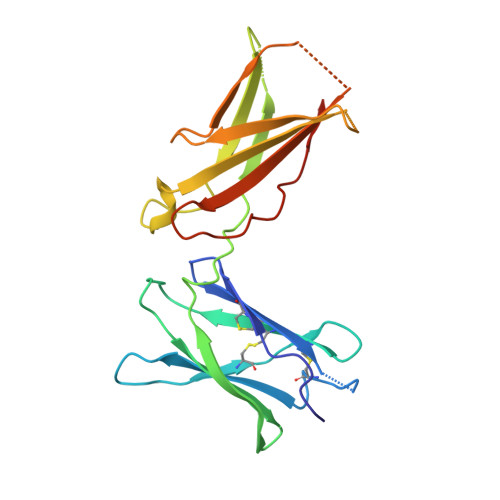
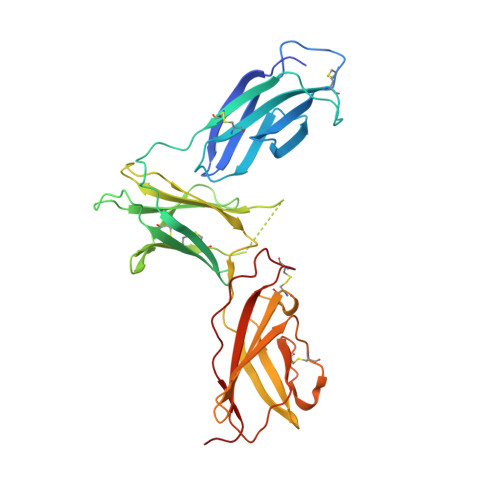
Instructive roles for cytokine-receptor binding parameters in determining signaling and functional potency.
Moraga, I., Richter, D., Wilmes, S., Winkelmann, H., Jude, K., Thomas, C., Suhoski, M.M., Engleman, E.G., Piehler, J., Garcia, K.C.(2015) Sci Signal 8: ra114-ra114
- PubMed: 26554818 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/scisignal.aab2677
- Primary Citation Related Structures:
5E4E - PubMed Abstract:
Cytokines dimerize cell surface receptors to activate signaling and regulate many facets of the immune response. Many cytokines have pleiotropic effects, inducing a spectrum of redundant and distinct effects on different cell types. This pleiotropy has hampered cytokine-based therapies, and the high doses required for treatment often lead to off-target effects, highlighting the need for a more detailed understanding of the parameters controlling cytokine-induced signaling and bioactivities. Using the prototypical cytokine interleukin-13 (IL-13), we explored the interrelationships between receptor binding and a wide range of downstream cellular responses. We applied structure-based engineering to generate IL-13 variants that covered a spectrum of binding strengths for the receptor subunit IL-13Rα1. Engineered IL-13 variants representing a broad range of affinities for the receptor exhibited similar potencies in stimulating the phosphorylation of STAT6 (signal transducer and activator of transcription 6). Delays in the phosphorylation and nuclear translocation of STAT6 were only apparent for those IL-13 variants with markedly reduced affinities for the receptor. From these data, we developed a mechanistic model that quantitatively reproduced the kinetics of STAT6 phosphorylation for the entire spectrum of binding affinities. Receptor endocytosis played a key role in modulating STAT6 activation, whereas the lifetime of receptor-ligand complexes at the plasma membrane determined the potency of the variant for inducing more distal responses. This complex interrelationship between extracellular ligand binding and receptor function provides the foundation for new mechanism-based strategies that determine the optimal cytokine dose to enhance therapeutic efficacy.
- Howard Hughes Medical Institute, Stanford University School of Medicine, Stanford, CA 94305-5345, USA. Department of Molecular and Cellular Physiology and Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305-5345, USA.
Organizational Affiliation: