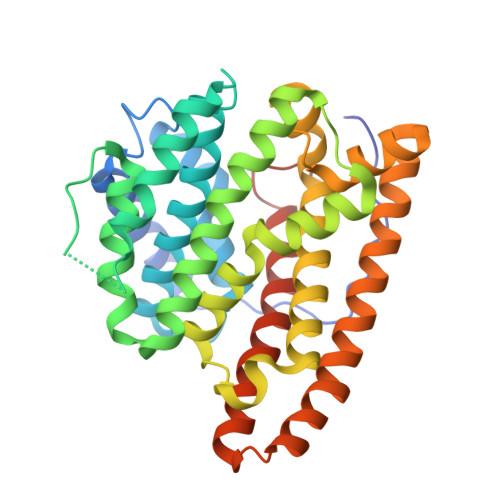
Structural Studies of Geosmin Synthase, a Bifunctional Sesquiterpene Synthase with alpha alpha Domain Architecture That Catalyzes a Unique Cyclization-Fragmentation Reaction Sequence.
Harris, G.G., Lombardi, P.M., Pemberton, T.A., Matsui, T., Weiss, T.M., Cole, K.E., Koksal, M., Murphy, F.V., Vedula, L.S., Chou, W.K., Cane, D.E., Christianson, D.W.(2015) Biochemistry 54: 7142-7155
- PubMed: 26598179 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.5b01143
- Primary Citation Related Structures:
5DW7, 5DZ2 - PubMed Abstract:
Geosmin synthase from Streptomyces coelicolor (ScGS) catalyzes an unusual, metal-dependent terpenoid cyclization and fragmentation reaction sequence. Two distinct active sites are required for catalysis: the N-terminal domain catalyzes the ionization and cyclization of farnesyl diphosphate to form germacradienol and inorganic pyrophosphate (PPi), and the C-terminal domain catalyzes the protonation, cyclization, and fragmentation of germacradienol to form geosmin and acetone through a retro-Prins reaction. A unique αα domain architecture is predicted for ScGS based on amino acid sequence: each domain contains the metal-binding motifs typical of a class I terpenoid cyclase, and each domain requires Mg(2+) for catalysis. Here, we report the X-ray crystal structure of the unliganded N-terminal domain of ScGS and the structure of its complex with three Mg(2+) ions and alendronate. These structures highlight conformational changes required for active site closure and catalysis. Although neither full-length ScGS nor constructs of the C-terminal domain could be crystallized, homology models of the C-terminal domain were constructed on the basis of ∼36% sequence identity with the N-terminal domain. Small-angle X-ray scattering experiments yield low-resolution molecular envelopes into which the N-terminal domain crystal structure and the C-terminal domain homology model were fit, suggesting possible αα domain architectures as frameworks for bifunctional catalysis.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania , Philadelphia, Pennsylvania 19104-6323, United States.
Organizational Affiliation: