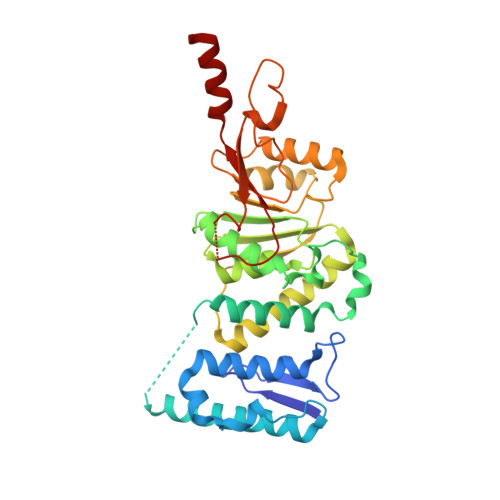
Optimization of a Fragment-Based Screening Hit toward Potent DOT1L Inhibitors Interacting in an Induced Binding Pocket.
Scheufler, C., Mobitz, H., Gaul, C., Ragot, C., Be, C., Fernandez, C., Beyer, K.S., Tiedt, R., Stauffer, F.(2016) ACS Med Chem Lett 7: 730-734
- PubMed: 27563394 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.6b00168
- Primary Citation Related Structures:
5DTM, 5DTQ, 5DTR - PubMed Abstract:
Mixed lineage leukemia (MLL) gene rearrangement induces leukemic transformation by ectopic recruitment of disruptor of telomeric silencing 1-like protein (DOT1L), a lysine histone methyltransferase, leading to local hypermethylation of H3K79 and misexpression of genes (including HoxA), which drive the leukemic phenotype. A weak fragment-based screening hit identified by SPR was cocrystallized with DOT1L and optimized using structure-based ligand optimization to yield compound 8 (IC50 = 14 nM). This series of inhibitors is structurally not related to cofactor SAM and is not interacting within the SAM binding pocket but induces a pocket adjacent to the SAM binding site.
- Novartis Institutes for Biomedical Research , 4002 Basel, Switzerland.
Organizational Affiliation: