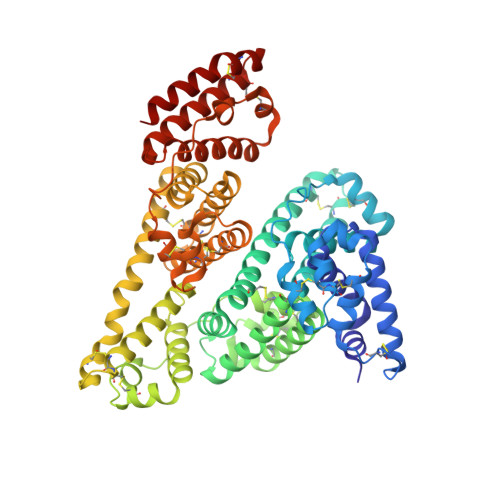
Crystal structure of equine serum albumin in complex with cetirizine reveals a novel drug binding site.
Handing, K.B., Shabalin, I.G., Szlachta, K., Majorek, K.A., Minor, W.(2016) Mol Immunol 71: 143-151
- PubMed: 26896718 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molimm.2016.02.003
- Primary Citation Related Structures:
5DQF - PubMed Abstract:
Serum albumin (SA) is the main transporter of drugs in mammalian blood plasma. Here, we report the first crystal structure of equine serum albumin (ESA) in complex with antihistamine drug cetirizine at a resolution of 2.1Å. Cetirizine is bound in two sites--a novel drug binding site (CBS1) and the fatty acid binding site 6 (CBS2). Both sites differ from those that have been proposed in multiple reports based on equilibrium dialysis and fluorescence studies for mammalian albumins as cetirizine binding sites. We show that the residues forming the binding pockets in ESA are highly conserved in human serum albumin (HSA), and suggest that binding of cetirizine to HSA will be similar. In support of that hypothesis, we show that the dissociation constants for cetirizine binding to CBS2 in ESA and HSA are identical using tryptophan fluorescence quenching. Presence of lysine and arginine residues that have been previously reported to undergo nonenzymatic glycosylation in CBS1 and CBS2 suggests that cetirizine transport in patients with diabetes could be altered. A review of all available SA structures from the PDB shows that in addition to the novel drug binding site we present here (CBS1), there are two pockets on SA capable of binding drugs that do not overlap with fatty acid binding sites and have not been discussed in published reviews.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, VA 22908-0736, USA; New York Structural Genomics Research Consortium (NYSGRC), USA.
Organizational Affiliation: