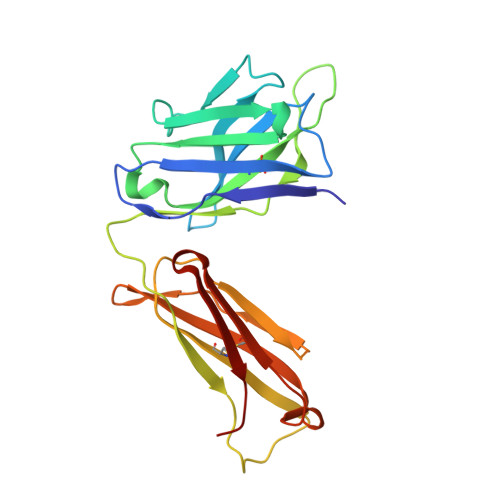
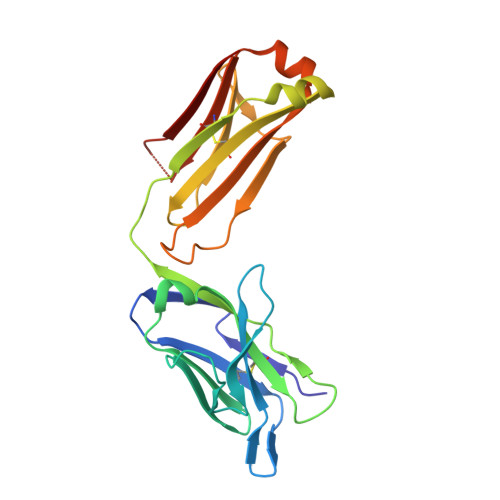
Lipid A-antibody structures reveal a widely-utilized pocket specific for negatively charged groups derived from from unrelated V-genes
Haji-Ghassemi, O., Muller-Loennies, S., Rodriguez, T., Brade, L., Kosma, P., Brade, H., Evans, S.V.(2015) J Biological Chem