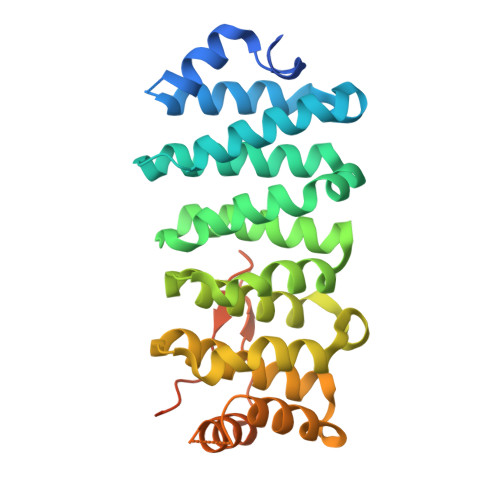
Crescerin uses a TOG domain array to regulate microtubules in the primary cilium.
Das, A., Dickinson, D.J., Wood, C.C., Goldstein, B., Slep, K.C.(2015) Mol Biol Cell 26: 4248-4264
- PubMed: 26378256 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1091/mbc.E15-08-0603
- Primary Citation Related Structures:
5DN7 - PubMed Abstract:
Eukaryotic cilia are cell-surface projections critical for sensing the extracellular environment. Defects in cilia structure and function result in a broad range of developmental and sensory disorders. However, mechanisms that regulate the microtubule (MT)-based scaffold forming the cilia core are poorly understood. TOG domain array-containing proteins ch-TOG and CLASP are key regulators of cytoplasmic MTs. Whether TOG array proteins also regulate ciliary MTs is unknown. Here we identify the conserved Crescerin protein family as a cilia-specific, TOG array-containing MT regulator. We present the crystal structure of mammalian Crescerin1 TOG2, revealing a canonical TOG fold with conserved tubulin-binding determinants. Crescerin1's TOG domains possess inherent MT-binding activity and promote MT polymerization in vitro. Using Cas9-triggered homologous recombination in Caenorhabditis elegans, we demonstrate that the worm Crescerin family member CHE-12 requires TOG domain-dependent tubulin-binding activity for sensory cilia development. Thus, Crescerin expands the TOG domain array-based MT regulatory paradigm beyond ch-TOG and CLASP, representing a distinct regulator of cilia structure.
- Department of Biochemistry and Biophysics, University of North Carolina, Chapel Hill, NC 27599 Molecular and Cellular Biophysics Program, University of North Carolina, Chapel Hill, NC 27599.
Organizational Affiliation: