Functional Antagonism of Human CD40 Achieved by Targeting a Unique Species-Specific Epitope.
Yamniuk, A.P., Suri, A., Krystek, S.R., Tamura, J., Ramamurthy, V., Kuhn, R., Carroll, K., Fleener, C., Ryseck, R., Cheng, L., An, Y., Drew, P., Grant, S., Suchard, S.J., Nadler, S.G., Bryson, J.W., Sheriff, S.(2016) J Mol Biology 428: 2860-2879
- PubMed: 27216500 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2016.05.014
- Primary Citation Related Structures:
5DMI, 5DMJ, 5IHL - PubMed Abstract:
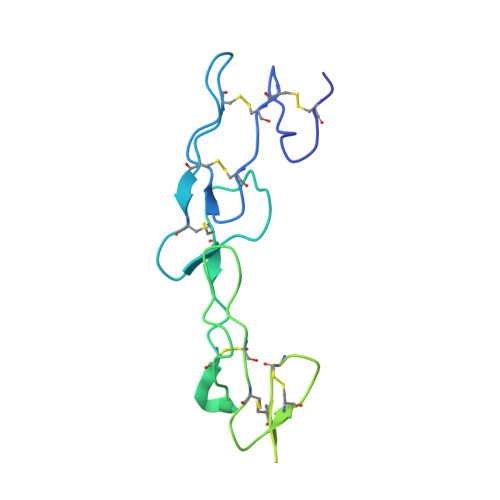
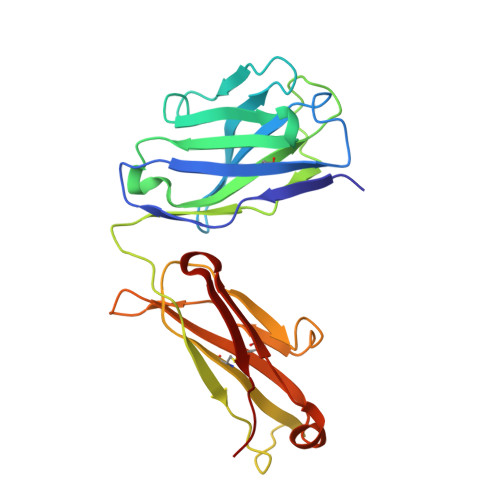
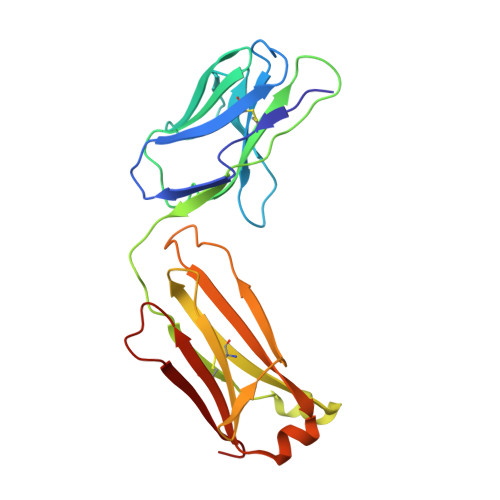
Current clinical anti-CD40 biologic agents include both antagonist molecules for the treatment of autoimmune diseases and agonist molecules for immuno-oncology, yet the relationship between CD40 epitope and these opposing biological outcomes is not well defined. This report describes the identification of potent antagonist domain antibodies (dAbs) that bind to a novel human CD40-specific epitope that is divergent in the CD40 of nonhuman primates. A similarly selected anti-cynomolgus CD40 dAb recognizing the homologous epitope is also a potent antagonist. Mutagenesis, biochemical, and X-ray crystallography studies demonstrate that the epitope is distinct from that of CD40 agonists. Both the human-specific and cynomolgus-specific molecules remain pure antagonists even when formatted as bivalent Fc-fusion proteins, making this an attractive therapeutic format for targeting hCD40 in autoimmune indications.
- Department of Molecular Discovery Technologies, Bristol-Myers Squibb, Princeton, NJ 08543, USA. Electronic address: aaron.yamniuk@bms.com.
Organizational Affiliation: