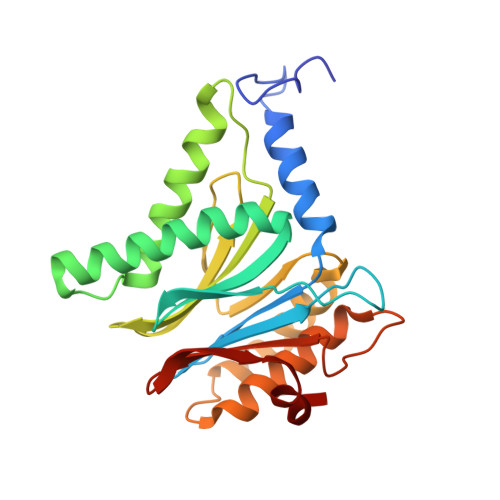
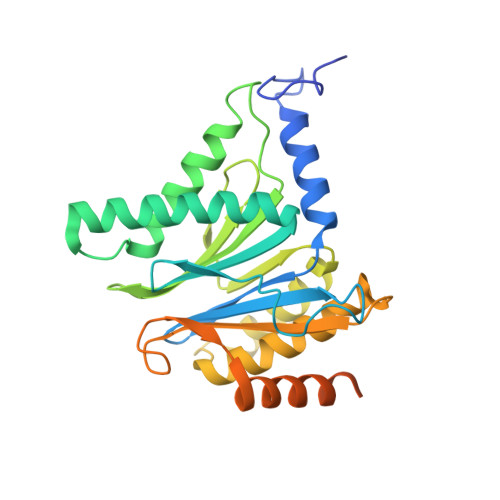
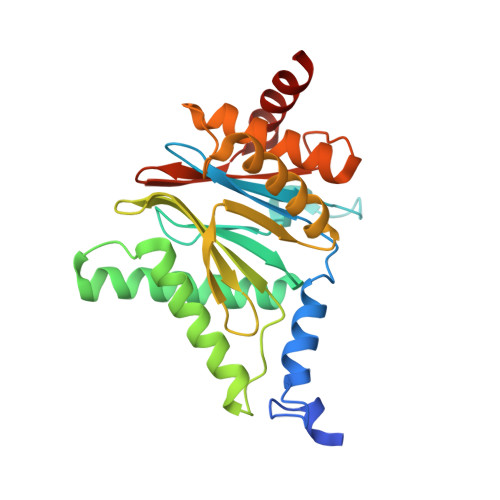
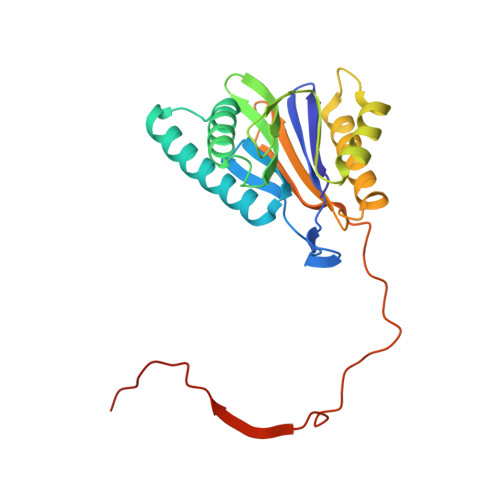
Targeted Delivery of Proteasome Inhibitors to Somatostatin-Receptor-Expressing Cancer Cells by Octreotide Conjugation.
Beck, P., Cui, H., Hegemann, J.D., Marahiel, M.A., Kruger, A., Groll, M.(2015) ChemMedChem 10: 1969-1973
- PubMed: 26471124 Search on PubMed
- DOI: https://doi.org/10.1002/cmdc.201500449
- Primary Citation Related Structures:
5DKI, 5DKJ - PubMed Abstract:
Clinical application of proteasome inhibitors (PIs) is so far limited to peripheral blood cancers due to the pronounced cytotoxicity towards all cell types. Targeted delivery of PIs could permit the treatment of other cancers along with decreasing side effects. Herein we describe the first small-molecule proteasome inhibitor conjugate for targeted delivery, created by fusing PIs to a synthetic ligand of somatostatin receptors, which are highly expressed in a variety of tumors. X-ray crystallographic studies and in vitro IC50 measurements demonstrated that addition of the cyclopeptide octreotide as a targeting vehicle does not affect the PI's binding mode. The cytotoxicity of the conjugate against somatostatin-receptor-expressing cells was up to 11-fold higher than that of a non-targeting surrogate. We have therefore established PIs as a new payload for drug conjugates and have shown that targeted delivery thereof could be a promising approach for the broader application of this FDA-approved class of compounds.
- Center for Integrated Protein Science Munich (CIPSM), Department of Chemistry, Technische Universität München, Lichtenbergstraße 4, 85748, Garching, Germany. pbeck@tum.de.
Organizational Affiliation: