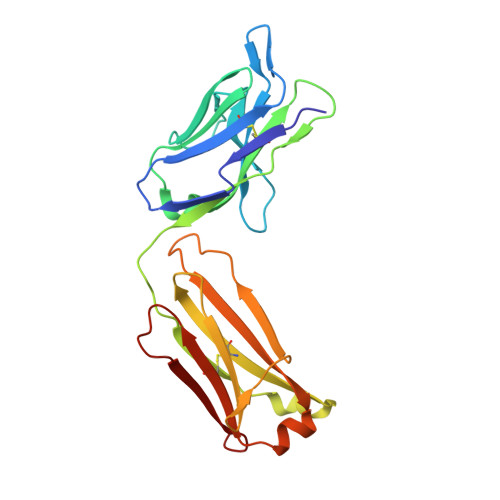
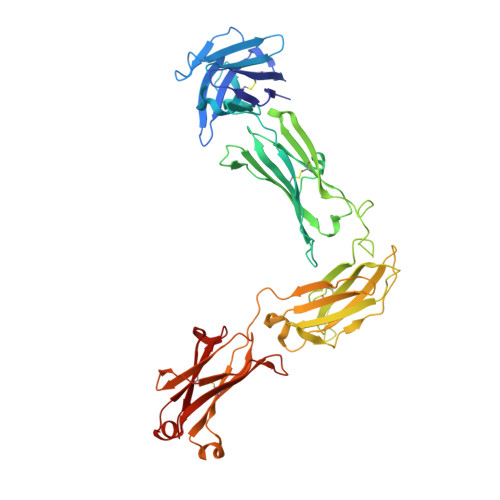
Structure of full-length human anti-PD1 therapeutic IgG4 antibody pembrolizumab.
Scapin, G., Yang, X., Prosise, W.W., McCoy, M., Reichert, P., Johnston, J.M., Kashi, R.S., Strickland, C.(2015) Nat Struct Mol Biol 22: 953-958
- PubMed: 26595420 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3129
- Primary Citation Related Structures:
5DK3 - PubMed Abstract:
Immunoglobulin G4 antibodies exhibit unusual properties with important biological consequences. We report the structure of the human full-length IgG4 S228P anti-PD1 antibody pembrolizumab, solved to 2.3-Å resolution. Pembrolizumab is a compact molecule, consistent with the presence of a short hinge region. The Fc domain is glycosylated at the CH2 domain on both chains, but one CH2 domain is rotated 120° with respect to the conformation observed in all reported structures to date, and its glycan chain faces the solvent. We speculate that this new conformation is driven by the shorter hinge. The structure suggests a role for the S228P mutation in preventing the IgG4 arm exchange. In addition, this unusual Fc conformation suggests possible structural diversity between IgG subclasses and shows that use of isolated antibody fragments could mask potentially important interactions, owing to molecular flexibility.
- Structural Chemistry, Merck &Co., Inc., Kenilworth, New Jersey, USA.
Organizational Affiliation: