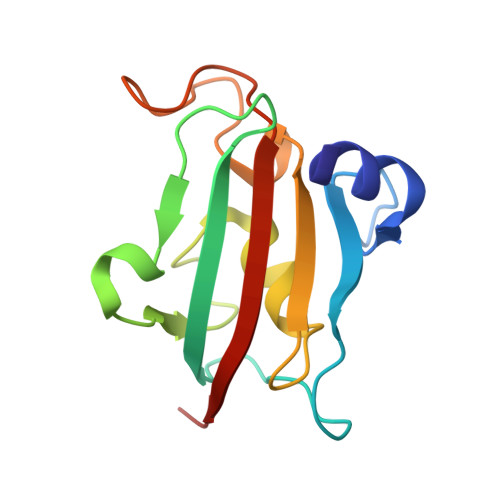
Structure-Affinity Relationship Analysis of Selective FKBP51 Ligands.
Feng, X., Sippel, C., Bracher, A., Hausch, F.(2015) J Med Chem 58: 7796-7806
- PubMed: 26419422 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00785
- Primary Citation Related Structures:
5DIT - PubMed Abstract:
The FK506-binding protein 51 (FKBP51) is a promising drug target for the treatment of stress-related psychiatric or metabolic disorders. Just recently, the first selective ligands for FKBP51 were reported based on an induced fit mechanism, but they are too large for a further drug development process. We therefore designed and synthesized a novel series of selective ligands to explore the requirements necessary for binding to the induced-fit conformation. All ligands of this series show no binding toward the structurally very similar antitarget FKBP52. With the cocrystal structure of the best ligand in this novel series we confirmed the induced fit mechanism. Furthermore, the structure-affinity relationship provides information about beneficial structural features, which is valuable for the development of improved FKBP51-directed drugs.
- Department of Translational Research in Psychiatry, Max Planck Institute of Psychiatry , Kraepelinstrasse 2, 80804 Munich, Germany.
Organizational Affiliation: