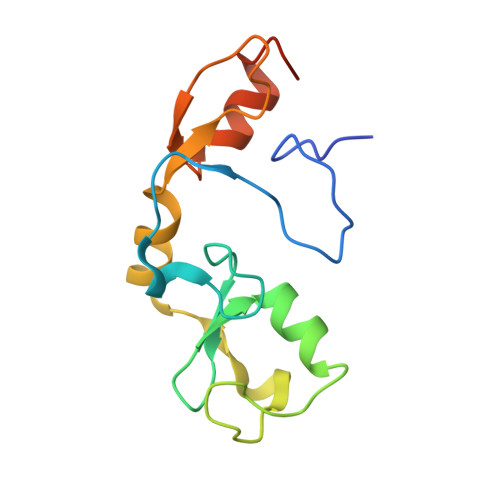
Structural basis for the indispensable role of a unique zinc finger motif in LNX2 ubiquitination.
Nayak, D., Sivaraman, J.(2015) Oncotarget 6: 34342-34357
- PubMed: 26451611 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.18632/oncotarget.5326
- Primary Citation Related Structures:
5DIN - PubMed Abstract:
LNX (Ligand of Numb Protein-X) proteins, LNX1 and LNX2, are RING- and PDZ-based E3-ubiquitin ligases known to interact with Numb. Silencing of LNX2 has been reported to down-regulate WNT and NOTCH, two key signaling pathways in tumorigenesis. Here we report the identification of the domain boundary of LNX2 to confer its ubiquitination activity, its crystal structure along with functional studies. We show that the RING domain in LNX2 is flanked by two Zinc-binding motifs (Zn-RING-Zn), in which the N-terminal Zinc-binding motif adopts novel conformation. Although this motif follows the typical Cys2His2-type zinc finger configuration, it is devoid of any secondary structure and forms an open circle conformation, which has not been reported yet. This unique N-terminal Zn-finger motif is indispensable for the activity and stability of LNX2, as verified using mutational studies. The Zn-RING-Zn domain of LNX2 is a dimer and assumes a rigid elongated structure that undergoes autoubiquitination and undergoes N-terminal polyubiquitination. The ubiquitin chains consist of all seven possible isopeptide linkages. These results were validated using full-length LNX2. Moreover we have demonstrated the ubiquitination of cell fate determinant protein, Numb by LNX2. Our study provides a structural basis for the functional machinery of LNX2 and thus provides the opportunity to investigate suitable drug targets against LNX2.
- Department of Biological Sciences, National University of Singapore, Singapore 117543.
Organizational Affiliation: