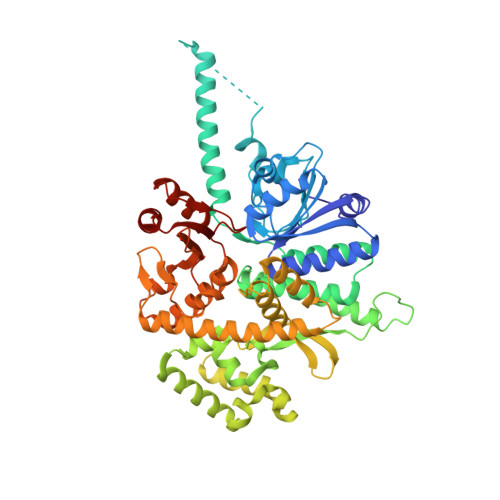
Structure of human Cdc45 and implications for CMG helicase function.
Simon, A.C., Sannino, V., Costanzo, V., Pellegrini, L.(2016) Nat Commun 7: 11638-11638
- PubMed: 27189187 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11638
- Primary Citation Related Structures:
5DGO - PubMed Abstract:
Cell division cycle protein 45 (Cdc45) is required for DNA synthesis during genome duplication, as a component of the Cdc45-MCM-GINS (CMG) helicase. Despite its essential biological function, its biochemical role in DNA replication has remained elusive. Here we report the 2.1-Å crystal structure of human Cdc45, which confirms its evolutionary link with the bacterial RecJ nuclease and reveals several unexpected features that underpin its function in eukaryotic DNA replication. These include a long-range interaction between N- and C-terminal DHH domains, blocking access to the DNA-binding groove of its RecJ-like fold, and a helical insertion in its N-terminal DHH domain, which appears poised for replisome interactions. In combination with available electron microscopy data, we validate by mutational analysis the mechanism of Cdc45 association with the MCM ring and GINS co-activator, critical for CMG assembly. These findings provide an indispensable molecular basis to rationalize the essential role of Cdc45 in genomic duplication.
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, UK.
Organizational Affiliation: