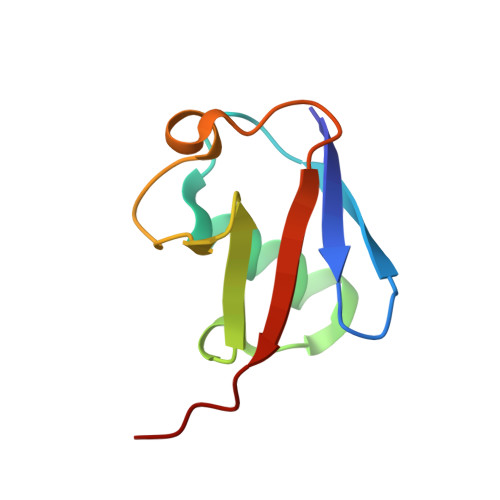
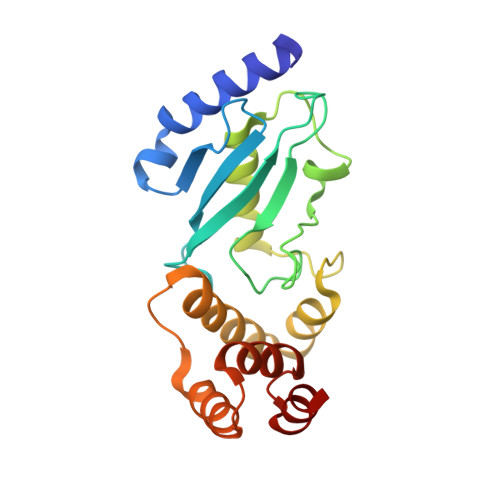
The molecular basis of lysine 48 ubiquitin chain synthesis by Ube2K.
Middleton, A.J., Day, C.L.(2015) Sci Rep 5: 16793-16793
- PubMed: 26592444 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep16793
- Primary Citation Related Structures:
5DFL - PubMed Abstract:
The post-translational modification of proteins by ubiquitin is central to the regulation of eukaryotic cells. Substrate-bound ubiquitin chains linked by lysine 11 and 48 target proteins to the proteasome for degradation and determine protein abundance in cells, while other ubiquitin chain linkages regulate protein interactions. The specificity of chain-linkage type is usually determined by ubiquitin-conjugating enzymes (E2s). The degradative E2, Ube2K, preferentially catalyses formation of Lys48-linked chains, but like most E2s, the molecular basis for chain formation is not well understood. Here we report the crystal structure of a Ube2K~ubiquitin conjugate and demonstrate that even though it is monomeric, Ube2K can synthesize Lys48-linked ubiquitin chains. Using site-directed mutagenesis and modelling, our studies reveal a molecular understanding of the catalytic complex and identify key features required for synthesis of degradative Lys48-linked chains. The position of the acceptor ubiquitin described here is likely conserved in other E2s that catalyse Lys48-linked ubiquitin chain synthesis.
- Department of Biochemistry, Otago School of Medical Sciences, University of Otago, Dunedin 9054, New Zealand.
Organizational Affiliation: