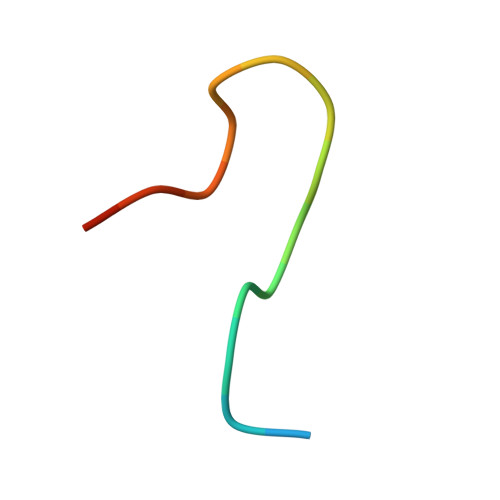
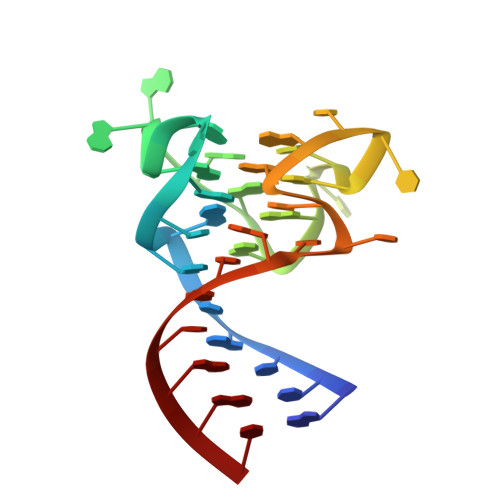
Crystal structure reveals specific recognition of a G-quadruplex RNA by a beta-turn in the RGG motif of FMRP.
Vasilyev, N., Polonskaia, A., Darnell, J.C., Darnell, R.B., Patel, D.J., Serganov, A.(2015) Proc Natl Acad Sci U S A 112: E5391-E5400
- PubMed: 26374839 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1515737112
- Primary Citation Related Structures:
5DE5, 5DE8, 5DEA - PubMed Abstract:
Fragile X Mental Retardation Protein (FMRP) is a regulatory RNA binding protein that plays a central role in the development of several human disorders including Fragile X Syndrome (FXS) and autism. FMRP uses an arginine-glycine-rich (RGG) motif for specific interactions with guanine (G)-quadruplexes, mRNA elements implicated in the disease-associated regulation of specific mRNAs. Here we report the 2.8-Å crystal structure of the complex between the human FMRP RGG peptide bound to the in vitro selected G-rich RNA. In this model system, the RNA adopts an intramolecular K(+)-stabilized G-quadruplex structure composed of three G-quartets and a mixed tetrad connected to an RNA duplex. The RGG peptide specifically binds to the duplex-quadruplex junction, the mixed tetrad, and the duplex region of the RNA through shape complementarity, cation-π interactions, and multiple hydrogen bonds. Many of these interactions critically depend on a type I β-turn, a secondary structure element whose formation was not previously recognized in the RGG motif of FMRP. RNA mutagenesis and footprinting experiments indicate that interactions of the peptide with the duplex-quadruplex junction and the duplex of RNA are equally important for affinity and specificity of the RGG-RNA complex formation. These results suggest that specific binding of cellular RNAs by FMRP may involve hydrogen bonding with RNA duplexes and that RNA duplex recognition can be a characteristic RNA binding feature for RGG motifs in other proteins.
- Department of Biochemistry and Molecular Pharmacology, New York University School of Medicine, New York, NY 10016;
Organizational Affiliation: