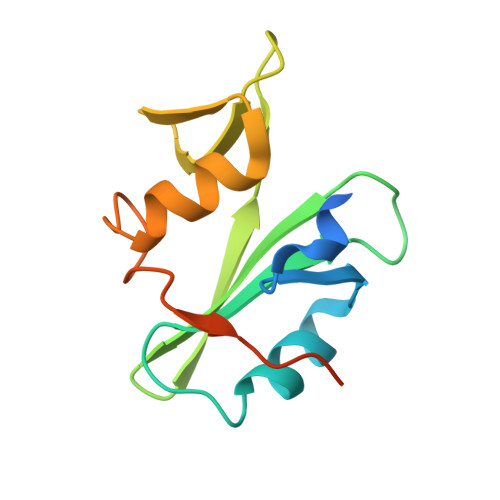
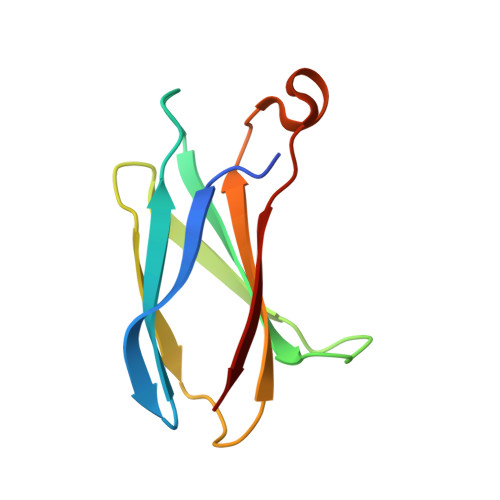
Allosteric Inhibition of Bcr-Abl Kinase by High Affinity Monobody Inhibitors Directed to the Src Homology 2 (SH2)-Kinase Interface.
Wojcik, J., Lamontanara, A.J., Grabe, G., Koide, A., Akin, L., Gerig, B., Hantschel, O., Koide, S.(2016) J Biological Chem 291: 8836-8847
- PubMed: 26912659 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.707901
- Primary Citation Related Structures:
5DC0, 5DC4, 5DC9 - PubMed Abstract:
Bcr-Abl is a constitutively active kinase that causes chronic myelogenous leukemia. We have shown that a tandem fusion of two designed binding proteins, termed monobodies, directed to the interaction interface between the Src homology 2 (SH2) and kinase domains and to the phosphotyrosine-binding site of the SH2 domain, respectively, inhibits the Bcr-Abl kinase activity. Because the latter monobody inhibits processive phosphorylation by Bcr-Abl and the SH2-kinase interface is occluded in the active kinase, it remained undetermined whether targeting the SH2-kinase interface alone was sufficient for Bcr-Abl inhibition. To address this question, we generated new, higher affinity monobodies with single nanomolar KD values targeting the kinase-binding surface of SH2. Structural and mutagenesis studies revealed the molecular underpinnings of the monobody-SH2 interactions. Importantly, the new monobodies inhibited Bcr-Abl kinase activity in vitro and in cells, and they potently induced cell death in chronic myelogenous leukemia cell lines. This work provides strong evidence for the SH2-kinase interface as a pharmacologically tractable site for allosteric inhibition of Bcr-Abl.
- From the Department of Biochemistry and Molecular Biology, The University of Chicago, Chicago, Illinois 60637.
Organizational Affiliation: