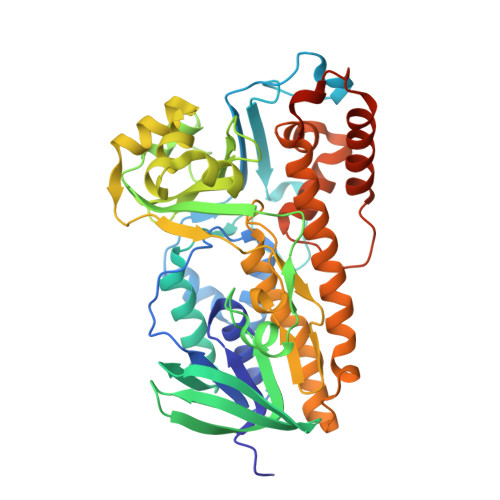
Crystal structure of halogenase PltA from the pyoluteorin biosynthetic pathway.
Pang, A.H., Garneau-Tsodikova, S., Tsodikov, O.V.(2015) J Struct Biol 192: 349-357
- PubMed: 26416533 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2015.09.013
- Primary Citation Related Structures:
5DBJ - PubMed Abstract:
Pyoluteorin is an antifungal agent composed of a 4,5-dichlorinated pyrrole group linked to a resorcinol moiety. The pyoluteorin biosynthetic gene cluster in Pseudomonas fluorescens Pf-5 encodes the halogenase PltA, which has been previously demonstrated to perform both chlorinations in vitro. PltA selectively accepts as a substrate a pyrrole moiety covalently tethered to a nonribosomal peptide thiolation domain PltL (pyrrolyl-S-PltL) for FAD-dependent di-chlorination, yielding 4,5-dichloropyrrolyl-S-PltL. We report a 2.75 Å-resolution crystal structure of PltA in complex with FAD and chloride. PltA is a dimeric enzyme, containing a flavin-binding fold conserved in flavin-dependent halogenases and monooxygenases, and an additional unique helical region at the C-terminus. This C-terminal region blocks a putative substrate-binding cleft, suggesting that a conformational change involving repositioning of this region is necessary to allow binding of the pyrrolyl-S-PltL substrate for its dichlorination by PltA.
- Department of Pharmaceutical Sciences, College of Pharmacy, University of Kentucky, 789 South Limestone Street, Lexington, KY 40536-0596, USA.
Organizational Affiliation: