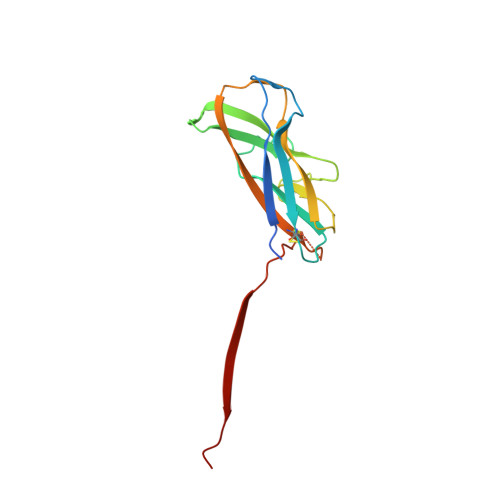
Crystal structure and analysis of HdaB: The enteroaggregative Escherichia coli AAF/IV pilus tip protein.
Lee, W.C., Matthews, S., Garnett, J.A.(2016) Protein Sci 25: 1898-1905
- PubMed: 27400770 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2982
- Primary Citation Related Structures:
5D55 - PubMed Abstract:
Enteroaggregative Escherichia coli is the primary cause of pediatric diarrhea in developing countries. They utilize aggregative adherence fimbriae (AAFs) to promote initial adherence to the host intestinal mucosa, promote the formation of biofilms, and mediate host invasion. Five AAFs have been identified to date and AAF/IV is amongst the most prevalent found in clinical isolates. Here we present the X-ray crystal structure of the AAF/IV tip protein HdaB at 2.0 Å resolution. It shares high structural homology with members of the Afa/Dr superfamily of fimbriae, which are involved in host invasion. We highlight surface exposed residues that share sequence homology and propose that these may function in invasion and also non-conserved regions that could mediate HdaB specific adhesive functions.
- Department of Life Sciences, Centre for Structural Biology, Imperial College London, South Kensington, London, SW7 2AZ, United Kingdom.
Organizational Affiliation: