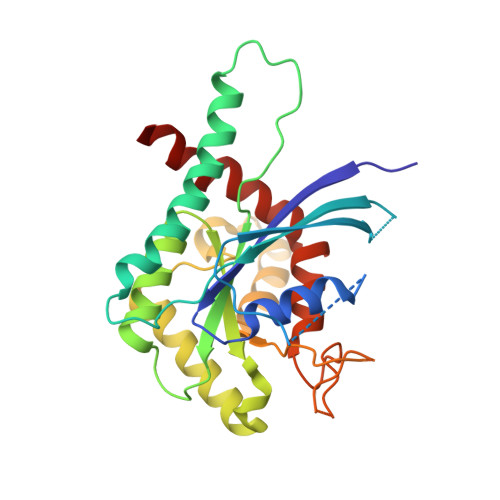
A complete compendium of crystal structures for the human SEPT3 subgroup reveals functional plasticity at a specific septin interface
Castro, D.K.S.V., da Silva, S.M.O., Pereira, H.M., Macedo, J.N.A., Leonardo, D.A., Valadares, N.F., Kumagai, P.S., Brandao-Neto, J., Araujo, A.P.U., Garratt, R.C.(2020) IUCrJ