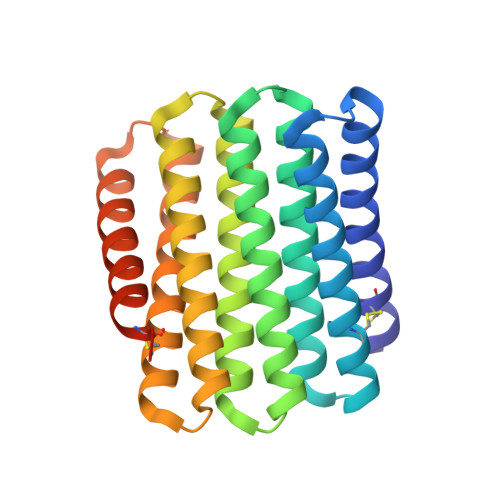
Exploring the repeat protein universe through computational protein design.
Brunette, T.J., Parmeggiani, F., Huang, P.S., Bhabha, G., Ekiert, D.C., Tsutakawa, S.E., Hura, G.L., Tainer, J.A., Baker, D.(2015) Nature 528: 580-584
- PubMed: 26675729 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature16162
- Primary Citation Related Structures:
5CWB, 5CWC, 5CWD, 5CWF, 5CWG, 5CWH, 5CWI, 5CWJ, 5CWK, 5CWL, 5CWM, 5CWN, 5CWO, 5CWP, 5CWQ - PubMed Abstract:
A central question in protein evolution is the extent to which naturally occurring proteins sample the space of folded structures accessible to the polypeptide chain. Repeat proteins composed of multiple tandem copies of a modular structure unit are widespread in nature and have critical roles in molecular recognition, signalling, and other essential biological processes. Naturally occurring repeat proteins have been re-engineered for molecular recognition and modular scaffolding applications. Here we use computational protein design to investigate the space of folded structures that can be generated by tandem repeating a simple helix-loop-helix-loop structural motif. Eighty-three designs with sequences unrelated to known repeat proteins were experimentally characterized. Of these, 53 are monomeric and stable at 95 °C, and 43 have solution X-ray scattering spectra consistent with the design models. Crystal structures of 15 designs spanning a broad range of curvatures are in close agreement with the design models with root mean square deviations ranging from 0.7 to 2.5 Å. Our results show that existing repeat proteins occupy only a small fraction of the possible repeat protein sequence and structure space and that it is possible to design novel repeat proteins with precisely specified geometries, opening up a wide array of new possibilities for biomolecular engineering.
- Department of Biochemistry, University of Washington, Seattle, Washington 98195, USA.
Organizational Affiliation: