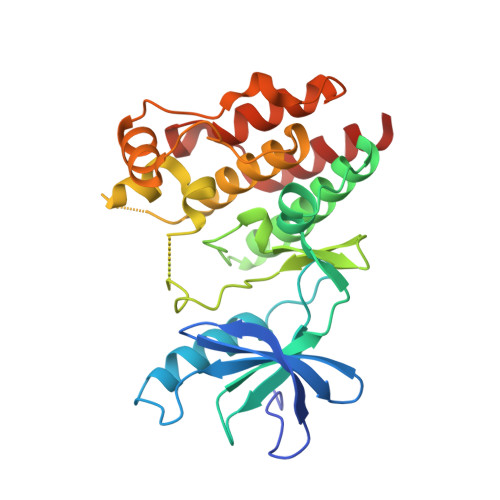
A Novel RAF Kinase Inhibitor with DFG-Out-Binding Mode: High Efficacy in BRAF-Mutant Tumor Xenograft Models in the Absence of Normal Tissue Hyperproliferation.
Waizenegger, I.C., Baum, A., Steurer, S., Stadtmuller, H., Bader, G., Schaaf, O., Garin-Chesa, P., Schlattl, A., Schweifer, N., Haslinger, C., Colbatzky, F., Mousa, S., Kalkuhl, A., Kraut, N., Adolf, G.R.(2016) Mol Cancer Ther 15: 354-365
- PubMed: 26916115 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-15-0617
- Primary Citation Related Structures:
5CSW, 5CSX - PubMed Abstract:
BI 882370 is a highly potent and selective RAF inhibitor that binds to the DFG-out (inactive) conformation of the BRAF kinase. The compound inhibited proliferation of human BRAF-mutant melanoma cells with 100× higher potency (1-10 nmol/L) than vemurafenib, whereas wild-type cells were not affected at 1,000 nmol/L. BI 882370 administered orally was efficacious in multiple mouse models of BRAF-mutant melanomas and colorectal carcinomas, and at 25 mg/kg twice daily showed superior efficacy compared with vemurafenib, dabrafenib, or trametinib (dosed to provide exposures reached in patients). To model drug resistance, A375 melanoma-bearing mice were initially treated with vemurafenib; all tumors responded with regression, but the majority subsequently resumed growth. Trametinib did not show any efficacy in this progressing population. BI 882370 induced tumor regression; however, resistance developed within 3 weeks. BI 882370 in combination with trametinib resulted in more pronounced regressions, and resistance was not observed during 5 weeks of second-line therapy. Importantly, mice treated with BI 882370 did not show any body weight loss or clinical signs of intolerability, and no pathologic changes were observed in several major organs investigated, including skin. Furthermore, a pilot study in rats (up to 60 mg/kg daily for 2 weeks) indicated lack of toxicity in terms of clinical chemistry, hematology, pathology, and toxicogenomics. Our results indicate the feasibility of developing novel compounds that provide an improved therapeutic window compared with first-generation BRAF inhibitors, resulting in more pronounced and long-lasting pathway suppression and thus improved efficacy.
- Department of Pharmacology and Translational Research, Boehringer Ingelheim RCV GmbH & Co KG, Vienna, Austria. irene.waizenegger@boehringer-ingelheim.com.
Organizational Affiliation: