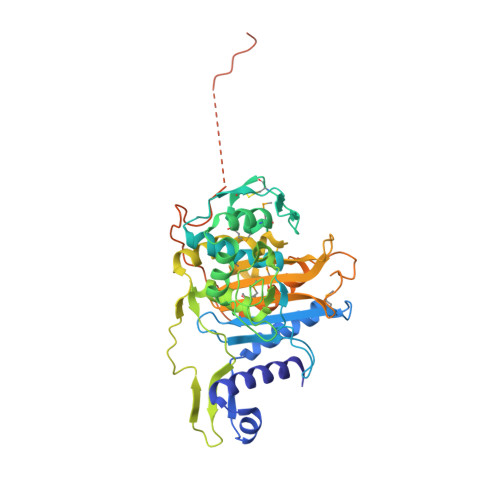
Crystal structures of the transpeptidase domain of the Mycobacterium tuberculosis penicillin-binding protein PonA1 reveal potential mechanisms of antibiotic resistance.
Filippova, E.V., Kieser, K.J., Luan, C.H., Wawrzak, Z., Kiryukhina, O., Rubin, E.J., Anderson, W.F.(2016) FEBS J 283: 2206-2218
- PubMed: 27101811 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.13738
- Primary Citation Related Structures:
5CRF, 5CXW - PubMed Abstract:
Mycobacterium tuberculosis is a human respiratory pathogen that causes the deadly disease tuberculosis. The rapid global spread of antibiotic-resistant M. tuberculosis makes tuberculosis infections difficult to treat. To overcome this problem new effective antimicrobial strategies are urgently needed. One promising target for new therapeutic approaches is PonA1, a class A penicillin-binding protein, which is required for maintaining physiological cell wall synthesis and cell shape during growth in mycobacteria. Here, crystal structures of the transpeptidase domain, the enzymatic domain responsible for penicillin binding, of PonA1 from M. tuberculosis in the inhibitor-free form and in complex with penicillin V are reported. We used site-directed mutagenesis, antibiotic profiling experiments, and fluorescence thermal shift assays to measure PonA1's sensitivity to different classes of β-lactams. Structural comparison of the PonA1 apo-form and the antibiotic-bound form shows that binding of penicillin V induces conformational changes in the position of the loop β4'-α3 surrounding the penicillin-binding site. We have also found that binding of different antibiotics including penicillin V positively impacts protein stability, while other tested β-lactams such as clavulanate or meropenem resulted in destabilization of PonA1. Our antibiotic profiling experiments indicate that the transpeptidase activity of PonA1 in both M. tuberculosis and M. smegmatis mediates tolerance to specific cell wall-targeting antibiotics, particularly to penicillin V and meropenem. Because M. tuberculosis is an important human pathogen, these structural data provide a template to design novel transpeptidase inhibitors to treat tuberculosis infections. Structural data are available in the PDB database under the accession numbers 5CRF and 5CXW.
- Department of Biochemistry and Molecular Genetics, Feinberg School of Medicine, Northwestern University, Chicago, IL, USA.
Organizational Affiliation: