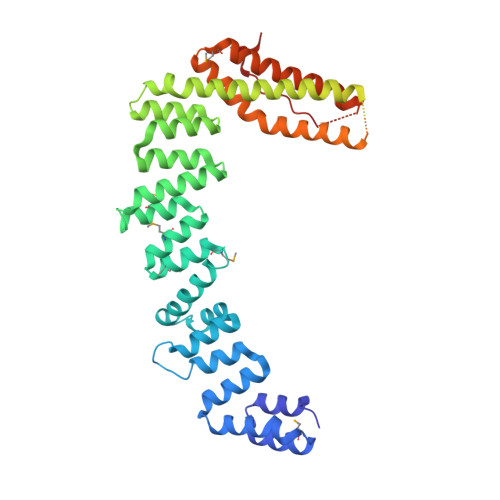
Dimerization of elongator protein 1 is essential for Elongator complex assembly.
Xu, H., Lin, Z., Li, F., Diao, W., Dong, C., Zhou, H., Xie, X., Wang, Z., Shen, Y., Long, J.(2015) Proc Natl Acad Sci U S A 112: 10697-10702
- PubMed: 26261306 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1502597112
- Primary Citation Related Structures:
5CQR, 5CQS - PubMed Abstract:
The evolutionarily conserved Elongator complex, which is composed of six subunits elongator protein 1 (Elp1 to -6), plays vital roles in gene regulation. The molecular hallmark of familial dysautonomia (FD) is the splicing mutation of Elp1 [also known as IκB kinase complex-associated protein (IKAP)] in the nervous system that is believed to be the primary cause of the devastating symptoms of this disease. Here, we demonstrate that disease-related mutations in Elp1 affect Elongator assembly, and we have determined the structure of the C-terminal portion of human Elp1 (Elp1-CT), which is sufficient for full-length Elp1 dimerization, as well as the structure of the cognate dimerization domain of yeast Elp1 (yElp1-DD). Our study reveals that the formation of the Elp1 dimer contributes to its stability in vitro and in vivo and is required for the assembly of both the human and yeast Elongator complexes. Functional studies suggest that Elp1 dimerization is essential for yeast viability. Collectively, our results identify the evolutionarily conserved dimerization domain of Elp1 and suggest that the pathological mechanisms underlying the onset and progression of Elp1 mutation-related disease may result from impaired Elongator activities.
- State Key Laboratory of Medicinal Chemical Biology, Nankai University, Tianjin 300071, China; College of Life Sciences, Nankai University, Tianjin 300071, China;
Organizational Affiliation: