Crystal structure of Ac-AChBP in complex with conotoxin GIC
Wang, X.Q., Xu, M.Y., Luo, S.L., Lin, B.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Soluble acetylcholine receptor | A, C [auth B], E [auth D], G, I | 236 | Aplysia californica | Mutation(s): 0 | 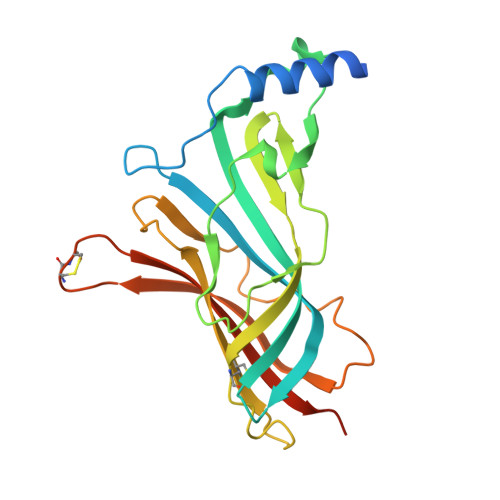 |
UniProt | |||||
Find proteins for Q8WSF8 (Aplysia californica) Explore Q8WSF8 Go to UniProtKB: Q8WSF8 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8WSF8 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Alpha-conotoxin GIC | B [auth F], D [auth C], F [auth E], H, J | 17 | Conus geographus | Mutation(s): 1 |  |
UniProt | |||||
Find proteins for Q86RB2 (Conus geographus) Explore Q86RB2 Go to UniProtKB: Q86RB2 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q86RB2 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 78.585 | α = 90 |
| b = 84.941 | β = 90 |
| c = 208.601 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |