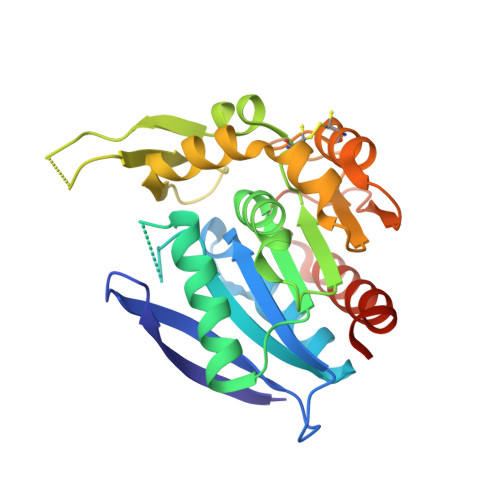
Characterisation of a New Family of Carboxyl Esterases with an OsmC Domain.
Jensen, M.V., Horsfall, L.E., Wardrope, C., Togneri, P.D., Marles-Wright, J., Rosser, S.J.(2016) PLoS One 11: e0166128-e0166128
- PubMed: 27851780 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0166128
- Primary Citation Related Structures:
5CML - PubMed Abstract:
Proteins in the serine esterase family are widely distributed in bacterial phyla and display activity against a range of biologically produced and chemically synthesized esters. A serine esterase from the psychrophilic bacterium Pseudoalteromonas arctica with a C-terminal OsmC-like domain was recently characterized; here we report on the identification and characterization of further putative esterases with OsmC-like domains constituting a new esterase family that is found in a variety of bacterial species from different environmental niches. All of these proteins contained the Ser-Asp-His motif common to serine esterases and a highly conserved pentapeptide nucleophilic elbow motif. We produced these proteins heterologously in Escherichia coli and demonstrated their activity against a range of esterase substrates. Two of the esterases characterized have activity of over two orders of magnitude higher than other members of the family, and are active over a wide temperature range. We determined the crystal structure of the esterase domain of the protein from Rhodothermus marinus and show that it conforms to the classical α/β hydrolase fold with an extended 'lid' region, which occludes the active site of the protein in the crystal. The expansion of characterized members of the esterase family and demonstration of activity over a wide-range of temperatures could be of use in biotechnological applications such as the pharmaceutical, detergent, bioremediation and dairy industries.
- Institute of Quantitative Biology, Biochemistry, and Biotechnology, School of Biological Sciences, University of Edinburgh, Edinburgh, United Kingdom.
Organizational Affiliation: