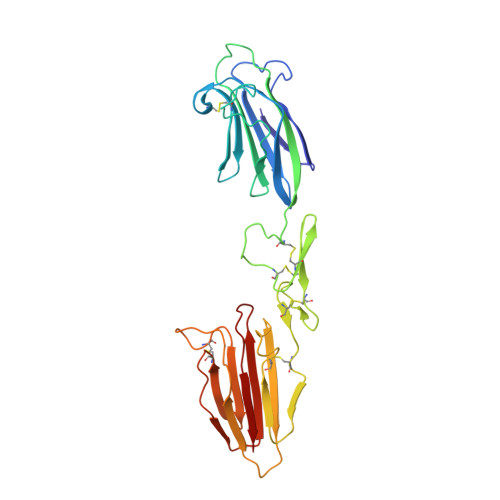
Flexibility in Mannan-Binding Lectin-Associated Serine Proteases-1 and -2 Provides Insight on Lectin Pathway Activation.
Nan, R., Furze, C.M., Wright, D.W., Gor, J., Wallis, R., Perkins, S.J.(2017) Structure 25: 364-375
- PubMed: 28111019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.12.014
- Primary Citation Related Structures:
5CIS, 5CKM, 5CKN, 5CKQ - PubMed Abstract:
The lectin pathway of complement is activated by complexes comprising a recognition component (mannose-binding lectin, serum ficolins, collectin-LK or collectin-K1) and a serine protease (MASP-1 or MASP-2). MASP-1 activates MASP-2, and MASP-2 cleaves C4 and C4b-bound C2. To clarify activation, new crystal structures of Ca 2+ -bound MASP dimers were determined, together with their solution structures from X-ray scattering, analytical ultracentrifugation, and atomistic modeling. Solution structures of the CUB1-EGF-CUB2 dimer of each MASP indicate that the two CUB2 domains were tilted by as much as 90° compared with the crystal structures, indicating considerable flexibility at the EGF-CUB2 junction. Solution structures of the full-length MASP dimers in their zymogen and activated forms revealed similar structures that were much more bent than anticipated from crystal structures. We conclude that MASP-1 and MASP-2 are flexible at multiple sites and that this flexibility may permit both intra- and inter-complex activation.
- Department of Structural and Molecular Biology, Division of Biosciences, University College London, Darwin Building, Gower Street, London WC1E 6BT, UK.
Organizational Affiliation: