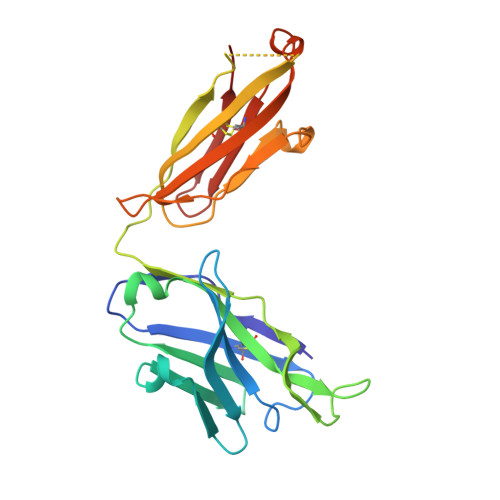
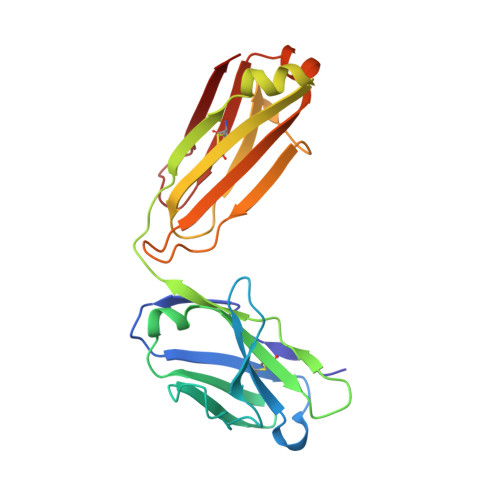
Structural and Thermodynamic Basis of Epitope Binding by Neutralizing and Nonneutralizing Forms of the Anti-HIV-1 Antibody 4E10
Rujas, E., Gulzar, N., Morante, K., Tsumoto, K., Scott, J.K., Nieva, J.L., Caaveiro, J.M.M.(2015) J Virol 89: 11975-11989
- PubMed: 26378169 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01793-15
- Primary Citation Related Structures:
5CIL, 5CIN, 5CIP - PubMed Abstract:
The 4E10 antibody recognizes the membrane-proximal external region (MPER) of the HIV-1 Env glycoprotein gp41 transmembrane subunit, exhibiting one of the broadest neutralizing activities known to date. The neutralizing activity of 4E10 requires solvent-exposed hydrophobic residues at the apex of the complementarity-determining region (CDR) H3 loop, but the molecular basis for this requirement has not been clarified. Here, we report the cocrystal structures and the energetic parameters of binding of a peptide bearing the 4E10-epitope sequence (4E10ep) to nonneutralizing versions of the 4E10 Fab. Nonneutralizing Fabs were obtained by shortening and decreasing the hydrophobicity of the CDR-H3 loop (termed ΔLoop) or by substituting the two tryptophan residues of the CDR-H3 apex with Asp residues (termed WDWD), which also decreases hydrophobicity but preserves the length of the loop. The analysis was complemented by the first crystal structure of the 4E10 Fab in its ligand-free state. Collectively, the data ruled out major conformational changes of CDR-H3 at any stage during the binding process (equilibrium or transition state). Although these mutations did not impact the affinity of wild-type Fab for the 4E10ep in solution, the two nonneutralizing versions of 4E10 were deficient in binding to MPER inserted in the plasma membrane (mimicking the environment faced by the antibody in vivo). The conclusions of our structure-function analysis strengthen the idea that to exert effective neutralization, the hydrophobic apex of the solvent-exposed CDR-H3 loop must recognize an antigenic structure more complex than just the linear α-helical epitope and likely constrained by the viral membrane lipids. The broadly neutralizing anti-HIV-1 4E10 antibody blocks infection caused by nearly all viral strains and isolates examined thus far. However, 4E10 (or 4E10-like) antibodies are rarely found in HIV-1-infected individuals or elicited through vaccination. Impediments to the design of successful 4E10 immunogens are partly attributed to an incomplete understanding of the structural and binding characteristics of this class of antibodies. Since the broadly neutralizing activity of 4E10 is abrogated by mutations of the tip of the CDR-H3, we investigated their impact on binding of the MPER-epitope at the atomic and energetic levels. We conclude that the difference between neutralizing and nonneutralizing antibodies of 4E10 is neither structural nor energetic but is related to the capacity to recognize the HIV-1 gp41 epitope inserted in biological membranes. Our findings strengthen the idea that to elicit similar neutralizing antibodies, the suitable MPER vaccine must be "delivered" in a membrane environment.
- Department of Bioengineering, Graduate School of Engineering, The University of Tokyo, Tokyo, Japan Biophysics Unit (CSIC, UPV/EHU) and Department of Biochemistry and Molecular Biology, University of the Basque Country (UPV/EHU), Bilbao, Spain.
Organizational Affiliation: