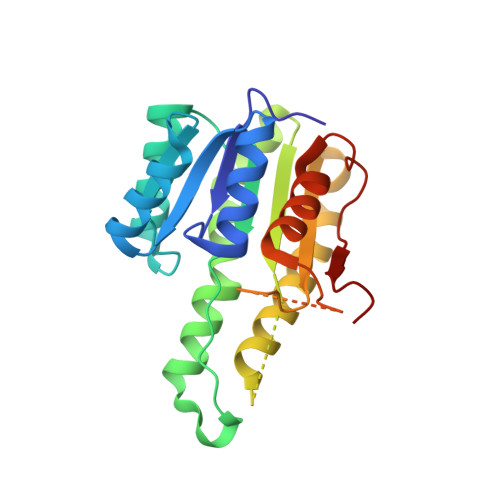
Structural Characterisation of FabG from Yersinia pestis, a Key Component of Bacterial Fatty Acid Synthesis.
Nanson, J.D., Forwood, J.K.(2015) PLoS One 10: e0141543-e0141543
- PubMed: 26539719 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0141543
- Primary Citation Related Structures:
5CDY, 5CEJ - PubMed Abstract:
Ketoacyl-acyl carrier protein reductases (FabG) are ubiquitously expressed enzymes that catalyse the reduction of acyl carrier protein (ACP) linked thioesters within the bacterial type II fatty acid synthesis (FASII) pathway. The products of these enzymes, saturated and unsaturated fatty acids, are essential components of the bacterial cell envelope. The FASII reductase enoyl-ACP reductase (FabI) has been the focus of numerous drug discovery efforts, some of which have led to clinical trials, yet few studies have focused on FabG. Like FabI, FabG appears to be essential for survival in many bacteria, similarly indicating the potential of this enzyme as a drug target. FabG enzymes are members of the short-chain alcohol dehydrogenase/reductase (SDR) family, and like other SDRs, exhibit highly conserved secondary and tertiary structures, and contain a number of conserved sequence motifs. Here we describe the crystal structures of FabG from Yersinia pestis (YpFabG), the causative agent of bubonic, pneumonic, and septicaemic plague, and three human pandemics. Y. pestis remains endemic in many parts of North America, South America, Southeast Asia, and Africa, and a threat to human health. YpFabG shares a high degree of structural similarity with bacterial homologues, and the ketoreductase domain of the mammalian fatty acid synthase from both Homo sapiens and Sus scrofa. Structural characterisation of YpFabG, and comparison with other bacterial FabGs and the mammalian fatty acid synthase, provides a strong platform for virtual screening of potential inhibitors, rational drug design, and the development of new antimicrobial agents to combat Y. pestis infections.
- School of Biomedical Sciences, Charles Sturt University, Wagga Wagga, NSW, 2678, Australia.
Organizational Affiliation: