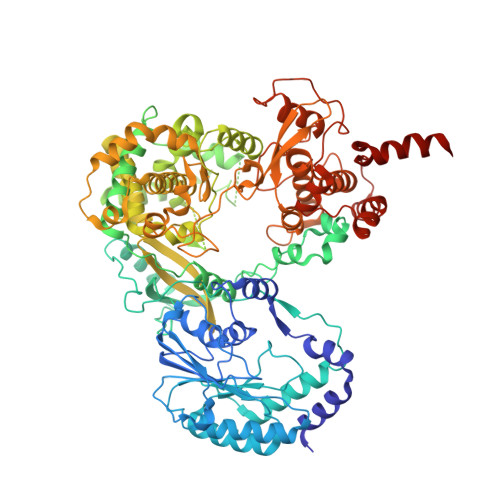
Dengue Virus Nonstructural Protein 5 (NS5) Assembles into a Dimer with a Unique Methyltransferase and Polymerase Interface.
Klema, V.J., Ye, M., Hindupur, A., Teramoto, T., Gottipati, K., Padmanabhan, R., Choi, K.H.(2016) PLoS Pathog 12: e1005451-e1005451
- PubMed: 26895240 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1005451
- Primary Citation Related Structures:
5CCV - PubMed Abstract:
Flavivirus nonstructural protein 5 (NS5) consists of methyltransferase (MTase) and RNA-dependent RNA polymerase (RdRp) domains, which catalyze 5'-RNA capping/methylation and RNA synthesis, respectively, during viral genome replication. Although the crystal structure of flavivirus NS5 is known, no data about the quaternary organization of the functional enzyme are available. We report the crystal structure of dengue virus full-length NS5, where eight molecules of NS5 are arranged as four independent dimers in the crystallographic asymmetric unit. The relative orientation of each monomer within the dimer, as well as the orientations of the MTase and RdRp domains within each monomer, is conserved, suggesting that these structural arrangements represent the biologically relevant conformation and assembly of this multi-functional enzyme. Essential interactions between MTase and RdRp domains are maintained in the NS5 dimer via inter-molecular interactions, providing evidence that flavivirus NS5 can adopt multiple conformations while preserving necessary interactions between the MTase and RdRp domains. Furthermore, many NS5 residues that reduce viral replication are located at either the inter-domain interface within a monomer or at the inter-molecular interface within the dimer. Hence the X-ray structure of NS5 presented here suggests that MTase and RdRp activities could be coordinated as a dimer during viral genome replication.
- Department of Biochemistry and Molecular Biology, Sealy Center for Structural Biology and Molecular Biophysics, University of Texas Medical Branch at Galveston, Galveston, Texas, United States of America.
Organizational Affiliation: