Combinatorial protein engineering of proteolytically resistant mesotrypsin inhibitors as candidates for cancer therapy.
Cohen, I., Kayode, O., Hockla, A., Sankaran, B., Radisky, D.C., Radisky, E.S., Papo, N.(2016) Biochem J 473: 1329-1341
- PubMed: 26957636 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20151410
- Primary Citation Related Structures:
5C67 - PubMed Abstract:
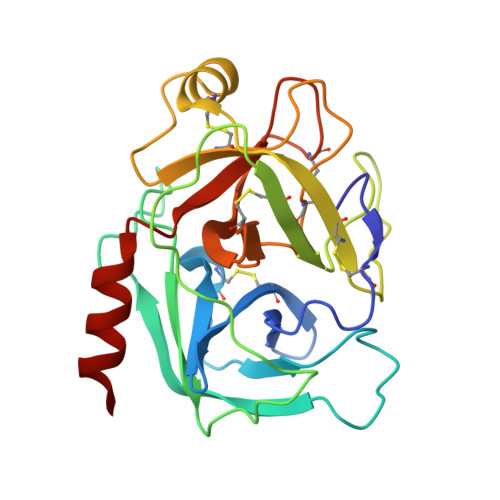
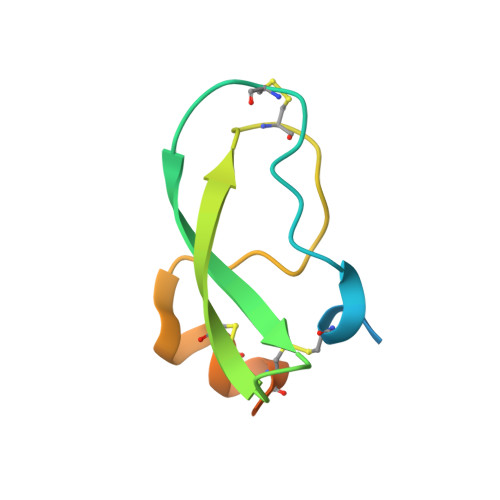
Engineered protein therapeutics offer advantages, including strong target affinity, selectivity and low toxicity, but like natural proteins can be susceptible to proteolytic degradation, thereby limiting their effectiveness. A compelling therapeutic target is mesotrypsin, a protease up-regulated with tumour progression, associated with poor prognosis, and implicated in tumour growth and progression of many cancers. However, with its unique capability for cleavage and inactivation of proteinaceous inhibitors, mesotrypsin presents a formidable challenge to the development of biological inhibitors. We used a powerful yeast display platform for directed evolution, employing a novel multi-modal library screening strategy, to engineer the human amyloid precursor protein Kunitz protease inhibitor domain (APPI) simultaneously for increased proteolytic stability, stronger binding affinity and improved selectivity for mesotrypsin inhibition. We identified a triple mutant APPIM17G/I18F/F34V, with a mesotrypsin inhibition constant (Ki) of 89 pM, as the strongest mesotrypsin inhibitor yet reported; this variant displays 1459-fold improved affinity, up to 350 000-fold greater specificity and 83-fold improved proteolytic stability compared with wild-type APPI. We demonstrated that APPIM17G/I18F/F34V acts as a functional inhibitor in cell-based models of mesotrypsin-dependent prostate cancer cellular invasiveness. Additionally, by solving the crystal structure of the APPIM17G/I18F/F34V-mesotrypsin complex, we obtained new insights into the structural and mechanistic basis for improved binding and proteolytic resistance. Our study identifies a promising mesotrypsin inhibitor as a starting point for development of anticancer protein therapeutics and establishes proof-of-principle for a novel library screening approach that will be widely applicable for simultaneously evolving proteolytic stability in tandem with desired functionality for diverse protein scaffolds.
- Department of Biotechnology Engineering and the National Institute of Biotechnology in the Negev, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel.
Organizational Affiliation: